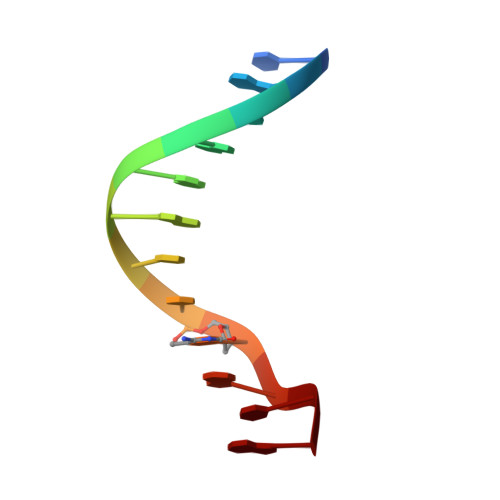
Structural insights into how 5-hydroxymethylation influences transcription factor binding.
Lercher, L., McDonough, M.A., El-Sagheer, A.H., Thalhammer, A., Kriaucionis, S., Brown, T., Schofield, C.J.(2014) Chem Commun (Camb) 50: 1794-1796
- PubMed: 24287551 Search on PubMed
- DOI: https://doi.org/10.1039/c3cc48151d
- Primary Citation Related Structures:
4C5X, 4C63, 4C64 - PubMed Abstract:
Transcription factor binding and high resolution crystallographic studies (1.3 Å) of Dickerson-Drew duplexes with cytosine, methylcytosine and hydroxymethylcytosine bases provide evidence that C-5 cytosine modifications could regulate transcription by context dependent effects on DNA transcription factor interactions.
- Department of Chemistry and the Oxford Centre for Integrative Systems Biology, Chemistry Research Laboratory, Oxford, UK. Christopher.schofield@chem.ox.ac.uk.
Organizational Affiliation: