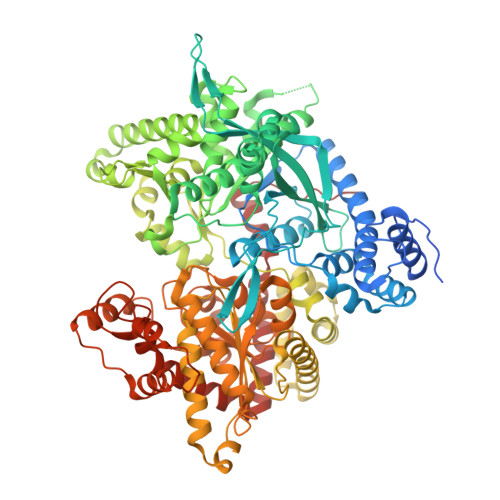
Sugar-Coated Sensor Chip and Nanoparticle Surfaces for the in Vitro Enzymatic Synthesis of Starch-Like Materials
O'Neill, E.C., Rashid, A.M., Stevenson, C.E.M., Hetru, A.C., Gunning, A.P., Rejzek, M., Nepogodiev, S.A., Bornemann, S., Lawson, D.M., Field, R.A.(2014) Chem Sci 5: 341