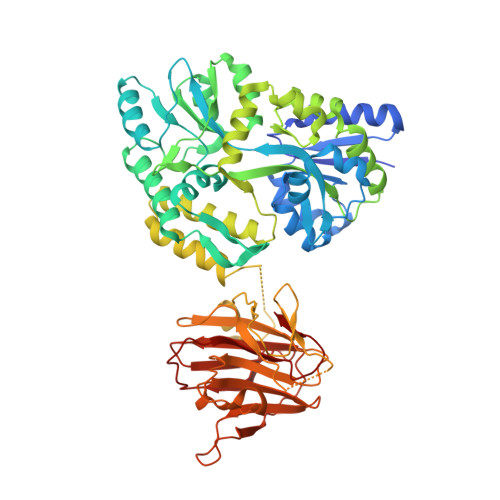
Structural Insight Into HIV-1 Capsid Recognition by Rhesus Trim5Alpha
Yang, H., Ji, X., Zhao, G., Ning, J., Zhao, Q., Aiken, C., Gronenborn, A.M., Zhang, P., Xiong, Y.(2012) Proc Natl Acad Sci U S A 109: 18372
- PubMed: 23091002 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.1210903109
- Primary Citation Related Structures:
4B3N - PubMed Abstract:
Tripartite motif protein isoform 5 alpha (TRIM5α) is a potent antiviral protein that restricts infection by HIV-1 and other retroviruses. TRIM5α recognizes the lattice of the retrovirus capsid through its B30.2 (PRY/SPRY) domain in a species-specific manner. Upon binding, TRIM5α induces premature disassembly of the viral capsid and activates the downstream innate immune response. We have determined the crystal structure of the rhesus TRIM5α PRY/SPRY domain that reveals essential features for capsid binding. Combined cryo-electron microscopy and biochemical data show that the monomeric rhesus TRIM5α PRY/SPRY, but not the human TRIM5α PRY/SPRY, can bind to HIV-1 capsid protein assemblies without causing disruption of the capsid. This suggests that the PRY/SPRY domain alone constitutes an important pattern-sensing component of TRIM5α that is capable of interacting with viral capsids of different curvatures. Our results provide molecular insights into the mechanisms of TRIM5α-mediated retroviral restriction.
- Department of Molecular Biophysics and Biochemistry, Yale University, New Haven, CT 06520, USA.
Organizational Affiliation: