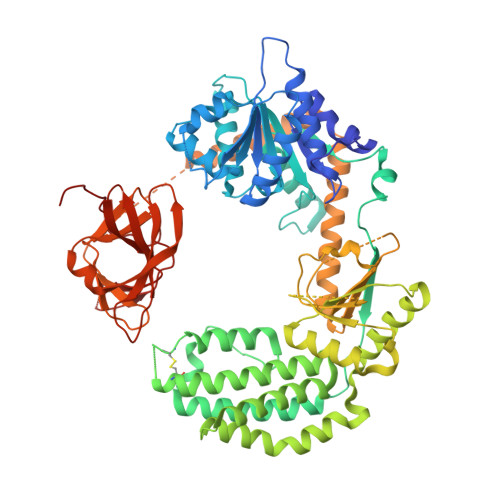
Defining the Functional Determinants for RNA Surveillance by Rig-I.
Kohlway, A., Luo, D., Rawling, D.C., Ding, S.C., Pyle, A.M.(2013) EMBO Rep 14: 772
- PubMed: 23897087 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/embor.2013.108
- Primary Citation Related Structures:
3ZD6, 3ZD7 - PubMed Abstract:
Retinoic acid-inducible gene-I (RIG-I) is an intracellular RNA sensor that activates the innate immune machinery in response to infection by RNA viruses. Here, we report the crystal structure of distinct conformations of a RIG-I:dsRNA complex, which shows that HEL2i-mediated scanning allows RIG-I to sense the length of RNA targets. To understand the implications of HEL2i scanning for catalytic activity and signalling by RIG-I, we examined its ATPase activity when stimulated by duplex RNAs of varying lengths and 5' composition. We identified a minimal RNA duplex that binds one RIG-I molecule, stimulates robust ATPase activity, and elicits a RIG-I-mediated interferon response in cells. Our results reveal that the minimal functional unit of the RIG-I:RNA complex is a monomer that binds at the terminus of a duplex RNA substrate. This behaviour is markedly different from the RIG-I paralog melanoma differentiation-associated gene 5 (MDA5), which forms cooperative filaments.
- Department of Molecular Biophysics and Biochemistry.
Organizational Affiliation: