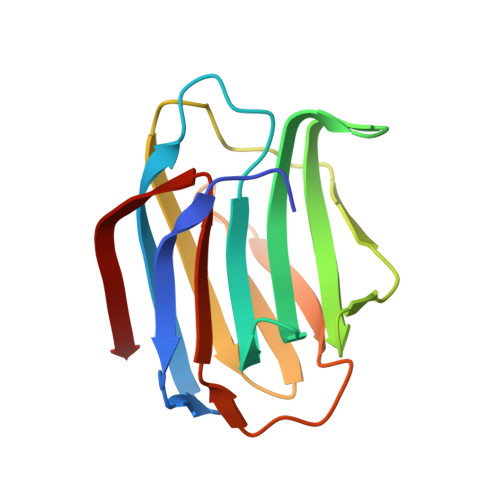
Crystal structure of a Xenopus laevis skin proto-type galectin, close to but distinct from galectin-1.
Nonaka, Y., Ogawa, T., Yoshida, H., Shoji, H., Nishi, N., Kamitori, S., Nakamura, T.(2015) Glycobiology 25: 792-803
- PubMed: 25804418 Search on PubMed
- DOI: https://doi.org/10.1093/glycob/cwv020
- Primary Citation Related Structures:
3WUC, 3WUD - PubMed Abstract:
Xenopus laevis (African clawed frog) has two types of proto-type galectins that are similar to mammalian galectin-1 in amino acid sequence. One type, comprising xgalectin-Ia and -Ib, is regarded as being equivalent to galectin-1, and the other type, comprising xgalectin-Va and -Vb, is expected to be a unique galectin subgroup. The latter is considerably abundant in frog skin; however, its biological function remains unclear. We determined the crystal structures of two proto-type galectins, xgalectin-Ib and -Va. The structures showed that both galectins formed a mammalian galectin-1-like homodimer, and furthermore, xgalectin-Va formed a homotetramer. This tetramer structure has not been reported for other galectins. Gel filtration and other experiments indicated that xgalectin-Va was in a dimer-tetramer equilibrium in solution, and lactose binding enhanced the tetramer formation. The residues involved in the dimer-dimer association were conserved in xgalectin-Va and -Vb, and one of the Xenopus (Silurana) tropicalis proto-type galectins, but not in xgalectin-Ia and -Ib, and other galectin-1-equivalent proteins. Xgalectin-Va preferred Galβ1-3GalNAc and not Galβ1-4GlcNAc, while xgalectin-Ib preferred Galβ1-4GlcNAc as well as human galectin-1. Xgalectin-Va/Vb would have diverged from the galectin-1 group with accompanying acquisition of the higher oligomer formation and altered ligand selectivity.
- Department of Endocrinology, Faculty of Medicine.
Organizational Affiliation: