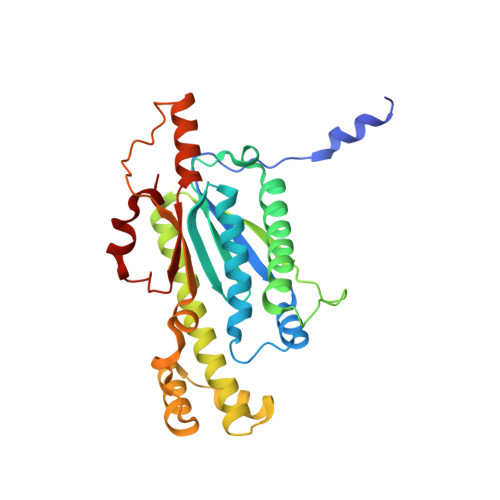
Structural basis of reverse nucleotide polymerization
Nakamura, A., Nemoto, T., Heinemann, I.U., Yamashita, K., Sonoda, T., Komoda, K., Tanaka, I., Soll, D., Yao, M.(2013) Proc Natl Acad Sci U S A 110: 20970-20975
- PubMed: 24324136 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.1321312111
- Primary Citation Related Structures:
3WBZ, 3WC0, 3WC1, 3WC2 - PubMed Abstract:
Nucleotide polymerization proceeds in the forward (5'-3') direction. This tenet of the central dogma of molecular biology is found in diverse processes including transcription, reverse transcription, DNA replication, and even in lagging strand synthesis where reverse polymerization (3'-5') would present a "simpler" solution. Interestingly, reverse (3'-5') nucleotide addition is catalyzed by the tRNA maturation enzyme tRNA(His) guanylyltransferase, a structural homolog of canonical forward polymerases. We present a Candida albicans tRNA(His) guanylyltransferase-tRNA(His) complex structure that reveals the structural basis of reverse polymerization. The directionality of nucleotide polymerization is determined by the orientation of approach of the nucleotide substrate. The tRNA substrate enters the enzyme's active site from the opposite direction (180° flip) compared with similar nucleotide substrates of canonical 5'-3' polymerases, and the finger domains are on opposing sides of the core palm domain. Structural, biochemical, and phylogenetic data indicate that reverse polymerization appeared early in evolution and resembles a mirror image of the forward process.
- Faculty of Advanced Life Science, Hokkaido University, Sapporo 060-0810, Japan.
Organizational Affiliation: