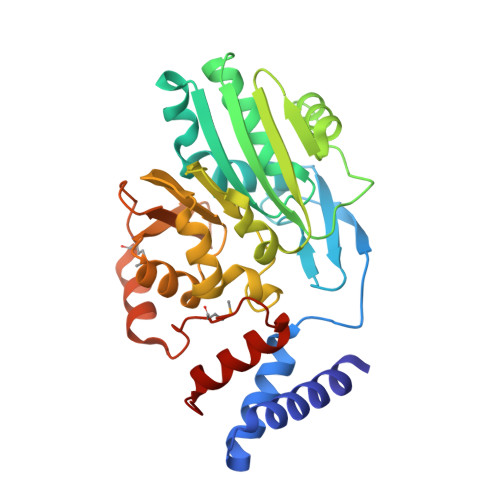
Characterization and structure of the Aquifex aeolicus protein DUF752: a bacterial tRNA-methyltransferase (MnmC2) functioning without the usually fused oxidase domain (MnmC1).
Kitamura, A., Nishimoto, M., Sengoku, T., Shibata, R., Jager, G., Bjork, G.R., Grosjean, H., Yokoyama, S., Bessho, Y.(2012) J Biological Chem 287: 43950-43960
- PubMed: 23091054 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M112.409300
- Primary Citation Related Structures:
3VYW - PubMed Abstract:
Post-transcriptional modifications of the wobble uridine (U34) of tRNAs play a critical role in reading NNA/G codons belonging to split codon boxes. In a subset of Escherichia coli tRNA, this wobble uridine is modified to 5-methylaminomethyluridine (mnm(5)U34) through sequential enzymatic reactions. Uridine 34 is first converted to 5-carboxymethylaminomethyluridine (cmnm(5)U34) by the MnmE-MnmG enzyme complex. The cmnm(5)U34 is further modified to mnm(5)U by the bifunctional MnmC protein. In the first reaction, the FAD-dependent oxidase domain (MnmC1) converts cmnm(5)U into 5-aminomethyluridine (nm(5)U34), and this reaction is immediately followed by the methylation of the free amino group into mnm(5)U34 by the S-adenosylmethionine-dependent domain (MnmC2). Aquifex aeolicus lacks a bifunctional MnmC protein fusion and instead encodes the Rossmann-fold protein DUF752, which is homologous to the methyltransferase MnmC2 domain of Escherichia coli MnmC (26% identity). Here, we determined the crystal structure of the A. aeolicus DUF752 protein at 2.5 Å resolution, which revealed that it catalyzes the S-adenosylmethionine-dependent methylation of nm(5)U in vitro, to form mnm(5)U34 in tRNA. We also showed that naturally occurring tRNA from A. aeolicus contains the 5-mnm group attached to the C5 atom of U34. Taken together, these results support the recent proposal of an alternative MnmC1-independent shortcut pathway for producing mnm(5)U34 in tRNAs.
- RIKEN SPring-8 Center, Harima Institute, 1-1-1 Kouto, Sayo, Hyogo 679-5148, Japan.
Organizational Affiliation: