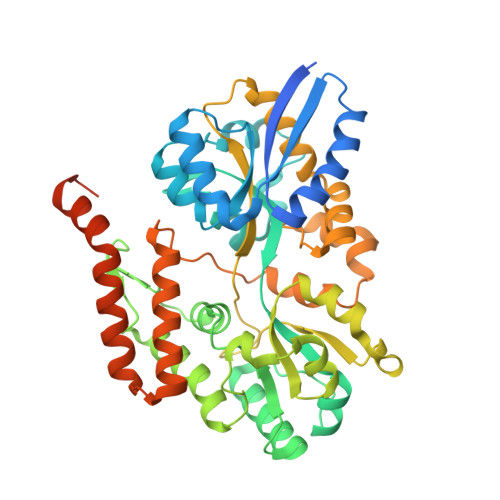
Structure determination of a sugar-binding protein from the phytopathogenic bacterium Xanthomonas citri.
Medrano, F.J., de Souza, C.S., Romero, A., Balan, A.(2014) Acta Crystallogr Sect F Struct Biol Cryst Commun 70: 564-571
- PubMed: 24817711 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S2053230X14006578
- Primary Citation Related Structures:
3UOR - PubMed Abstract:
The uptake of maltose and related sugars in Gram-negative bacteria is mediated by an ABC transporter encompassing a periplasmic component (the maltose-binding protein or MalE), a pore-forming membrane protein (MalF and MalG) and a membrane-associated ATPase (MalK). In the present study, the structure determination of the apo form of the putative maltose/trehalose-binding protein (Xac-MalE) from the citrus pathogen Xanthomonas citri in space group P6522 is described. The crystals contained two protein molecules in the asymmetric unit and diffracted to 2.8 Å resolution. Xac-MalE conserves the structural and functional features of sugar-binding proteins and a ligand-binding pocket with similar characteristics to eight different orthologues, including the residues for maltose and trehalose interaction. This is the first structure of a sugar-binding protein from a phytopathogenic bacterium, which is highly conserved in all species from the Xanthomonas genus.
- Department of Chemical and Physical Biology, Centro de Investigaciones Biologicas (CSIC), Madrid, Spain.
Organizational Affiliation: