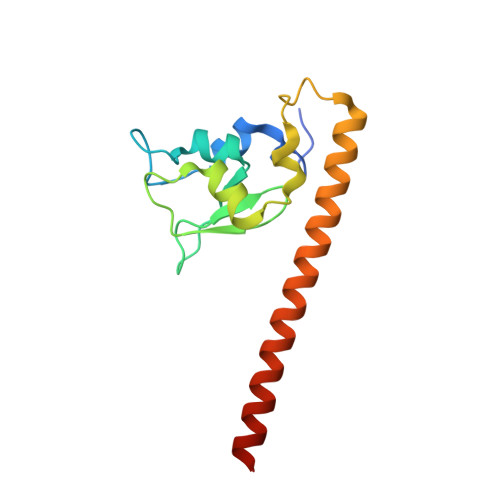
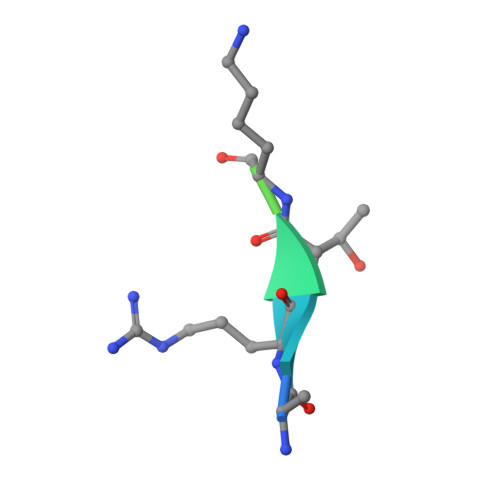
Structural Basis for Recognition of H3T3ph and Smac/DIABLO N-terminal Peptides by Human Survivin.
Du, J., Kelly, A.E., Funabiki, H., Patel, D.J.(2012) Structure 20: 185-195
- PubMed: 22244766 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.str.2011.12.001
- Primary Citation Related Structures:
3UIG, 3UIH, 3UII, 3UIJ, 3UIK - PubMed Abstract:
Survivin is an inhibitor of apoptosis family protein implicated in apoptosis and mitosis. In apoptosis, it has been shown to recognize the Smac/DIABLO protein. It is also a component of the chromosomal passenger complex, a key player during mitosis. Recently, Survivin was identified in vitro and in vivo as the direct binding partner for phosphorylated Thr3 on histone H3 (H3T3ph). We have undertaken structural and binding studies to investigate the molecular basis underlying recognition of H3T3ph and Smac/DIABLO N-terminal peptides by Survivin. Our crystallographic studies establish recognition of N-terminal Ala in both complexes and identify intermolecular hydrogen-bonding interactions in the Survivin phosphate-binding pocket that contribute to H3T3ph mark recognition. In addition, our calorimetric data establish that Survivin binds tighter to the H3T3ph-containing peptide relative to the N-terminal Smac/DIABLO peptide, and this preference can be reversed through structure-guided mutations that increase the hydrophobicity of the phosphate-binding pocket.
- Structural Biology Program, Memorial Sloan-Kettering Cancer Center, New York, NY 10065, USA.
Organizational Affiliation: