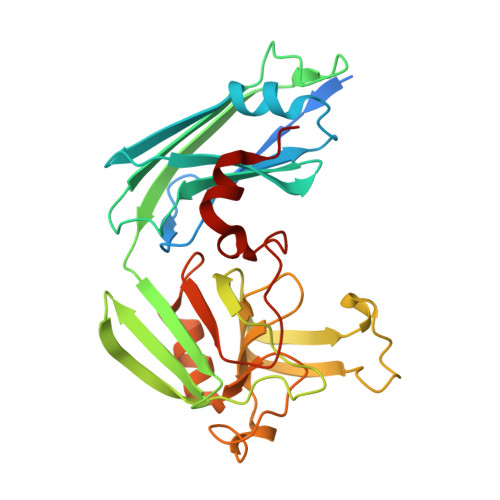
Targeting the Cell Wall of Mycobacterium tuberculosis: Structure and Mechanism of L,D-Transpeptidase 2.
Erdemli, S.B., Gupta, R., Bishai, W.R., Lamichhane, G., Amzel, L.M., Bianchet, M.A.(2012) Structure 20: 2103-2115
- PubMed: 23103390 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.str.2012.09.016
- Primary Citation Related Structures:
3TUR, 3TX4, 3U1P, 3VAE - PubMed Abstract:
With multidrug-resistant cases of tuberculosis increasing globally, better antibiotic drugs and novel drug targets are becoming an urgent need. Traditional β-lactam antibiotics that inhibit D,D-transpeptidases are not effective against mycobacteria, in part because mycobacteria rely mostly on L,D-transpeptidases for biosynthesis and maintenance of their peptidoglycan layer. This reliance plays a major role in drug resistance and persistence of Mycobacterium tuberculosis (Mtb) infections. The crystal structure at 1.7 Å resolution of the Mtb L,D-transpeptidase Ldt(Mt2) containing a bound peptidoglycan fragment, reported here, provides information about catalytic site organization as well as substrate recognition by the enzyme. Based on our structural, kinetic, and calorimetric data, we propose a catalytic mechanism for Ldt(Mt2) in which both acyl-acceptor and acyl-donor substrates reach the catalytic site from the same, rather than different, entrances. Together, this information provides vital insights to facilitate development of drugs targeting this validated yet unexploited enzyme.
- Department of Biophysics and Biophysical Chemistry, Johns Hopkins University School of Medicine, Baltimore, MD 21205, USA.
Organizational Affiliation: