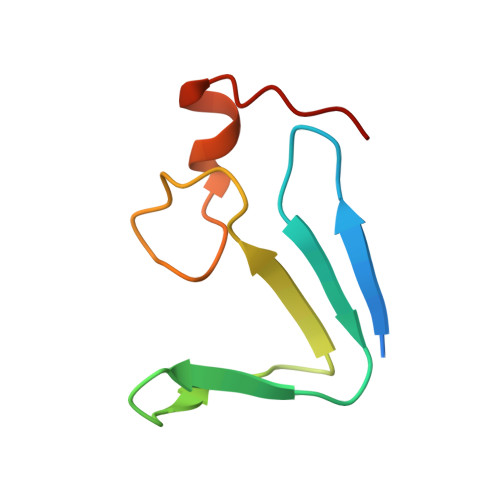
Structure and Molecular Evolution of CDGSH Iron-Sulfur Domains.
Lin, J., Zhang, L., Lai, S., Ye, K.(2011) PLoS One 6: e24790-e24790
- PubMed: 21949752 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1371/journal.pone.0024790
- Primary Citation Related Structures:
3TBM, 3TBN, 3TBO - PubMed Abstract:
The recently discovered CDGSH iron-sulfur domains (CISDs) are classified into seven major types with a wide distribution throughout the three domains of life. The type 1 protein mitoNEET has been shown to fold into a dimer with the signature CDGSH motif binding to a [2Fe-2S] cluster. However, the structures of all other types of CISDs were unknown. Here we report the crystal structures of type 3, 4, and 6 CISDs determined at 1.5 Å, 1.8 Å and 1.15 Å resolution, respectively. The type 3 and 4 CISD each contain one CDGSH motif and adopt a dimeric structure. Although similar to each other, the two structures have permutated topologies, and both are distinct from the type 1 structure. The type 6 CISD contains tandem CDGSH motifs and adopts a monomeric structure with an internal pseudo dyad symmetry. All currently known CISD structures share dual iron-sulfur binding modules and a β-sandwich for either intermolecular or intramolecular dimerization. The iron-sulfur binding module, the β-strand N-terminal to the module and a proline motif are conserved among different type structures, but the dimerization module and the interface and orientation between the two iron-sulfur binding modules are divergent. Sequence analysis further shows resemblance between CISD types 4 and 7 and between 1 and 2. Our findings suggest that all CISDs share common ancestry and diverged into three primary folds with a characteristic phylogenetic distribution: a eukaryote-specific fold adopted by types 1 and 2 proteins, a prokaryote-specific fold adopted by types 3, 4 and 7 proteins, and a tandem-motif fold adopted by types 5 and 6 proteins. Our comprehensive structural, sequential and phylogenetic analysis provides significant insight into the assembly principles and evolutionary relationship of CISDs.
- National Institute of Biological Sciences, Beijing, China.
Organizational Affiliation: