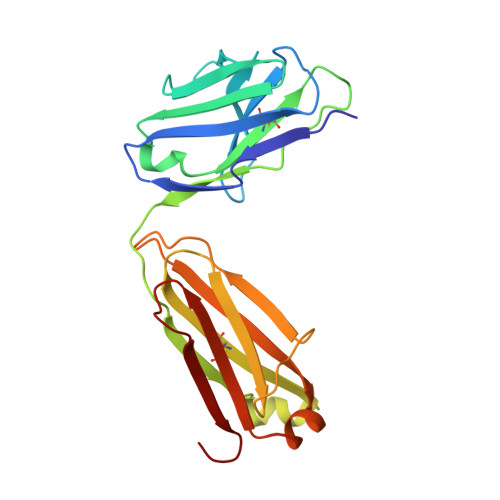
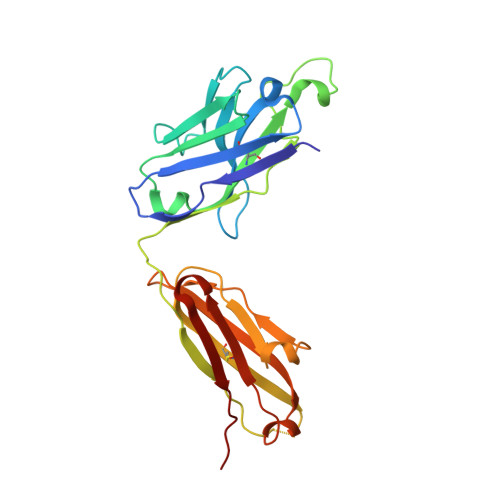
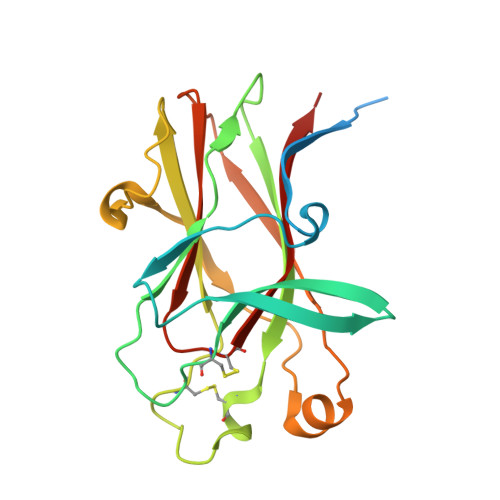
Structural and functional characterization of an agonistic anti-human EphA2 monoclonal antibody.
Peng, L., Oganesyan, V., Damschroder, M.M., Wu, H., Dall'Acqua, W.F.(2011) J Mol Biology 413: 390-405
- PubMed: 21867711 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2011.08.018
- Primary Citation Related Structures:
3SKJ - PubMed Abstract:
We report here the three-dimensional structure of human ephrin type A receptor 2 (EphA2) bound to the Fab (fragment antigen binding) of an agonistic human antibody (1C1; IgG1/κ). The structure of the corresponding complex was solved at a resolution of 2.5 Å using molecular replacement and constitutes the first reported structure of a human ephrin receptor bound to an antibody. We have also defined the corresponding functional epitope using a mutagenesis-based approach. This study revealed discrete structural features that determine the fine specificity of 1C1 to EphA2. Our data also provided a molecular basis for 1C1 mechanism of action. More precisely, we propose that its agonistic, internalizing properties are the result of ligand mimicry by the third heavy-chain complementarity-determining region of 1C1. Because EphA2 is an important contributor to cancer formation and progression, these findings may have implications for designing the next generation of anti-tumor therapies.
- Department of Antibody Discovery and Protein Engineering, MedImmune, Gaithersburg, MD 20878, USA.
Organizational Affiliation: