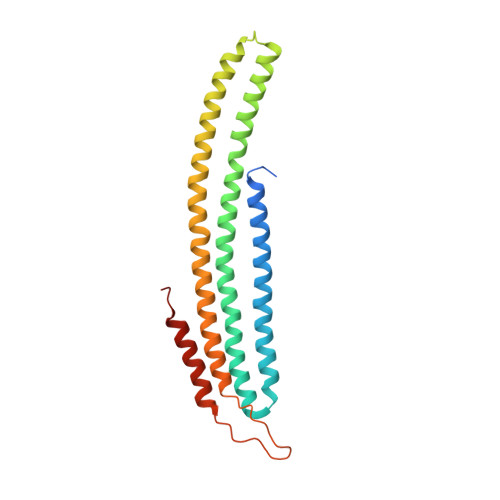
Eisosome-driven plasma membrane organization is mediated by BAR domains.
Ziolkowska, N.E., Karotki, L., Rehman, M., Huiskonen, J.T., Walther, T.C.(2011) Nat Struct Mol Biol 18: 854-856
- PubMed: 21685922 Search on PubMed
- DOI: https://doi.org/10.1038/nsmb.2080
- Primary Citation Related Structures:
3PLT - PubMed Abstract:
Plasma membranes are organized into domains of different protein and lipid composition. Eisosomes are key complexes for yeast plasma membrane organization, containing primarily Pil1 and Lsp1. Here we show that both proteins consist mostly of a banana-shaped BAR domain common to membrane sculpting proteins, most similar to the ones of amphiphysin, arfaptin 2 and endophilin 2. Our data reveal a previously unrecognized family of BAR-domain proteins involved in plasma membrane organization.
- Max Planck Institute of Biochemistry, Organelle Architecture and Dynamics, Martinsried, Germany.
Organizational Affiliation: