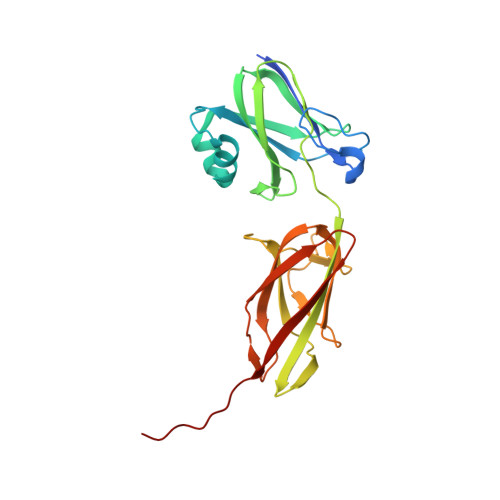
An IgG-like domain in the minor pilin GBS52 of Streptococcus agalactiae mediates lung epithelial cell adhesion.
Krishnan, V., Gaspar, A.H., Ye, N., Mandlik, A., Ton-That, H., Narayana, S.V.(2007) Structure 15: 893-903
- PubMed: 17697995 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.str.2007.06.015
- Primary Citation Related Structures:
3PHS - PubMed Abstract:
Streptococcus agalactiae is the leading cause of neonatal pneumonia, sepsis, and meningitis. The pathogen assembles heterotrimeric pilus structures on its surface; however, their function in pathogenesis is poorly understood. We report here the crystal structure of the pilin GBS52, which reveals two IgG-like fold domains, N1 and N2. Each domain is comprised of seven antiparallel beta strands, an arrangement similar to the fold observed in the Staphylococcus aureus adhesin Cna. Consistent with its role as an adhesin, deletion of gbs52 gene significantly reduces bacterial adherence to pulmonary epithelial cells. Moreover, latex beads linked to the GBS52 protein adhere to pulmonary but not to many other epithelial cells; binding to the former is specifically inhibited by antibodies against GBS52. Nonetheless, substantial binding is only observed with N2 domain-conjugated beads. This study presents the structure of a Gram-positive pilin that utilizes a distinct IgG fold variant to mediate pathogen adherence to a specific tissue.
- School of Optometry and Center for Biophysical Sciences and Engineering, University of Alabama at Birmingham, AL 35294, USA.
Organizational Affiliation: