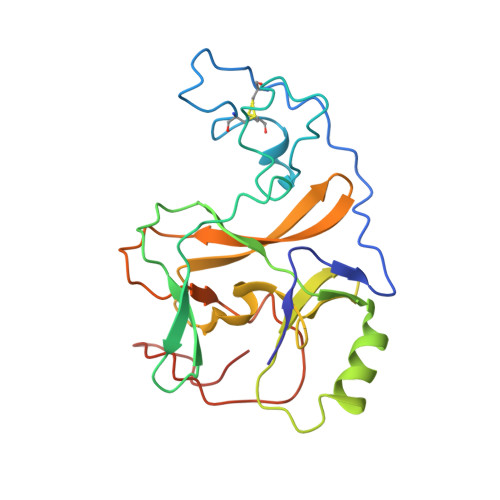
Crystal structure of the human histone methyltransferase ASH1L catalytic domain and its implications for the regulatory mechanism
An, S., Yeo, K.J., Jeon, Y.H., Song, J.(2011) J Biological Chem 286: 8369-8374
- PubMed: 21239497 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M110.203380
- Primary Citation Related Structures:
3OPE - PubMed Abstract:
Absent, small, or homeotic disc1 (Ash1) is a trithorax group histone methyltransferase that is involved in gene activation. Although there are many known histone methyltransferases, their regulatory mechanisms are poorly understood. Here, we present the crystal structure of the human ASH1L catalytic domain, showing its substrate binding pocket blocked by a loop from the post-SET domain. In this configuration, the loop limits substrate access to the active site. Mutagenesis of the loop stimulates ASH1L histone methyltransferase activity, suggesting that ASH1L activity may be regulated through the loop from the post-SET domain. In addition, we show that human ASH1L specifically methylates histone H3 Lys-36. Our data implicate that there may be a regulatory mechanism of ASH1L histone methyltransferases.
- From the Structural Biology Laboratory of Epigenetics, Department of Biological Sciences, Graduate school of Nanoscience and Technology (World Class University), KI for the BioCentury, Korea Advanced Institute of Science and Technology (KAIST), 335 Gwahangno, Yuseong-gu, Daejeon 305-701 and.
Organizational Affiliation: