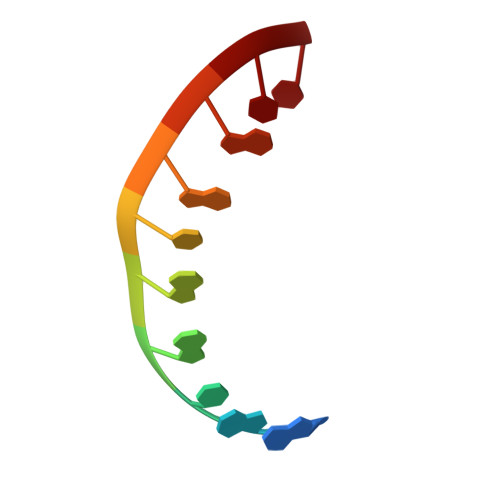
Atomic resolution structure of CAG RNA repeats: structural insights and implications for the trinucleotide repeat expansion diseases.
Kiliszek, A., Kierzek, R., Krzyzosiak, W.J., Rypniewski, W.(2010) Nucleic Acids Res 38: 8370-8376
- PubMed: 20702420 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1093/nar/gkq700
- Primary Citation Related Structures:
3NJ6, 3NJ7 - PubMed Abstract:
CAG repeats occur predominantly in the coding regions of human genes, which suggests their functional importance. In some genes, these sequences can undergo pathogenic expansions leading to neurodegenerative polyglutamine (poly-Q) diseases. The mutant transcripts containing expanded CAG repeats possibly contribute to pathogenesis in addition to the well-known pathogenic effects of mutant proteins. We have analysed two crystal forms of RNA duplexes containing CAG repeats: (GGCAGCAGCC)(2). One of the structures has been determined at atomic resolution (0.95 Å) and the other at 1.9 Å. The duplexes include non-canonical A-A pairs that fit remarkably well within a regular A-helix. All the adenosines are in the anti-conformation and the only interaction within each A-A pair is a single C2-H2···N1 hydrogen bond. Both adenosines in each A-A pair are shifted towards the major groove, although to different extents; the A which is the H-bond donor stands out more (the 'thumbs-up' conformation). The main effect on the helix conformation is a local unwinding. The CAG repeats and the previously examined CUG structures share a similar pattern of electrostatic charge distribution in the minor groove, which could explain their affinity for the pathogenesis-related MBNL1 protein.
- Institute of Bioorganic Chemistry, Polish Academy of Sciences, Noskowskiego 12/14, 61-704 Poznan, Poland.
Organizational Affiliation: