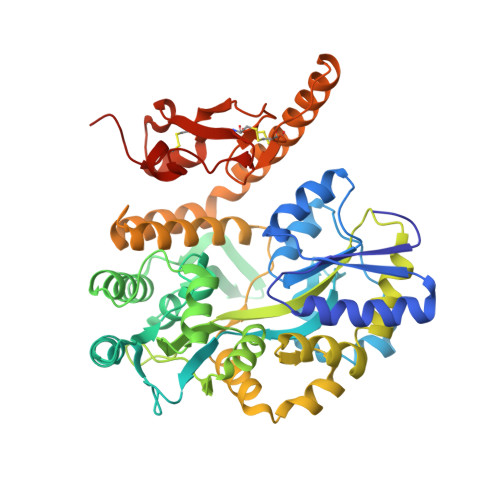
Crystal Structure of the PAC1R Extracellular Domain Unifies a Consensus Fold for Hormone Recognition by Class B G-Protein Coupled Receptors.
Kumar, S., Pioszak, A., Zhang, C., Swaminathan, K., Xu, H.E.(2011) PLoS One 6: e19682-e19682
- PubMed: 21625560 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1371/journal.pone.0019682
- Primary Citation Related Structures:
3N94 - PubMed Abstract:
Pituitary adenylate cyclase activating polypeptide (PACAP) is a member of the PACAP/glucagon family of peptide hormones, which controls many physiological functions in the immune, nervous, endocrine, and muscular systems. It activates adenylate cyclase by binding to its receptor, PAC1R, a member of class B G-protein coupled receptors (GPCR). Crystal structures of a number of Class B GPCR extracellular domains (ECD) bound to their respective peptide hormones have revealed a consensus mechanism of hormone binding. However, the mechanism of how PACAP binds to its receptor remains controversial as an NMR structure of the PAC1R ECD/PACAP complex reveals a different topology of the ECD and a distinct mode of ligand recognition. Here we report a 1.9 Å crystal structure of the PAC1R ECD, which adopts the same fold as commonly observed for other members of Class B GPCR. Binding studies and cell-based assays with alanine-scanned peptides and mutated receptor support a model that PAC1R uses the same conserved fold of Class B GPCR ECD for PACAP binding, thus unifying the consensus mechanism of hormone binding for this family of receptors.
- Laboratory of Structural Sciences, Van Andel Research Institute, Grand Rapids, Michigan, United States of America.
Organizational Affiliation: