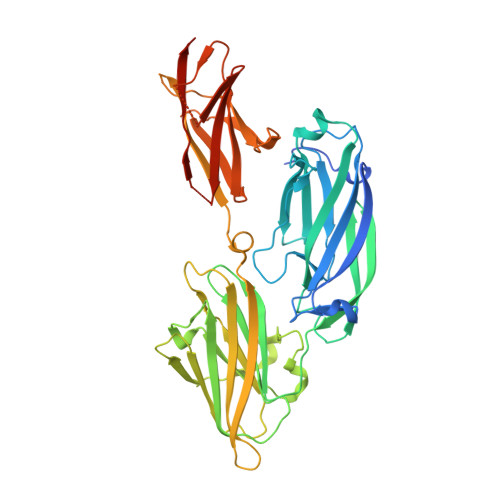
Crystal structure of the functional region of Uro-adherence factor A from Staphylococcus saprophyticus reveals participation of the B domain in ligand binding
Matsuoka, E., Tanaka, Y., Kuroda, M., Shouji, Y., Ohta, T., Tanaka, I., Yao, M.(2011) Protein Sci 20: 406-416
- PubMed: 21280131 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1002/pro.573
- Primary Citation Related Structures:
3IRP, 3IRZ, 3IS1 - PubMed Abstract:
Staphylococci use cell wall-anchored proteins as adhesins to attach to host tissues. Staphylococcus saprophyticus, a uropathogenic species, has a unique cell wall-anchored protein, uro-adherence factor A (UafA), which shows erythrocyte binding activity. To investigate the mechanism of adhesion by UafA, we determined the crystal structure of the functional region of UafA at 1.5 Å resolution. The structure was composed of three domains, designated as the N2, N3, and B domains, arranged in a triangular relative configuration. Hemagglutination inhibition assay with domain-truncated mutants indicated that both N and B domains were necessary for erythrocyte binding. Based on these results, a novel manner of ligand binding in which the B domain acts as a functional domain was proposed as the adhesion mechanism of S. saprophyticus.
- Graduate School of Life Science, Hokkaido University, Sapporo 060-0810, Japan.
Organizational Affiliation: