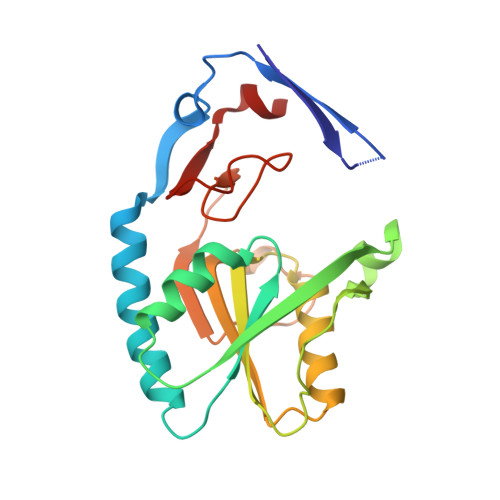
2.06 Angstrom resolution structure of a hypoxanthine-guanine phosphoribosyltransferase (hpt-1) from Bacillus anthracis str. 'Ames Ancestor'
Halavaty, A.S., Shuvalova, L., Minasov, G., Dubrovska, I., Peterson, S.N., Anderson, W.F., Center for Structural Genomics of Infectious Diseases (CSGID)To be published.