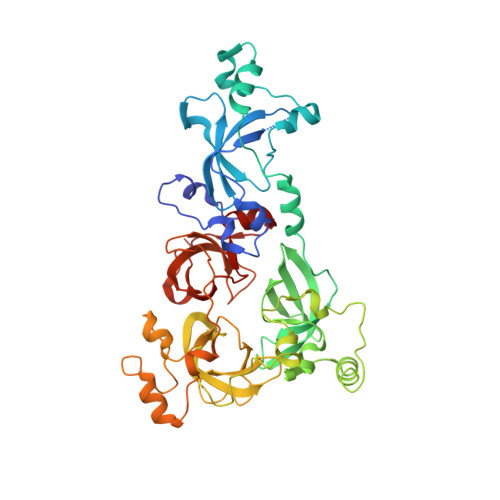
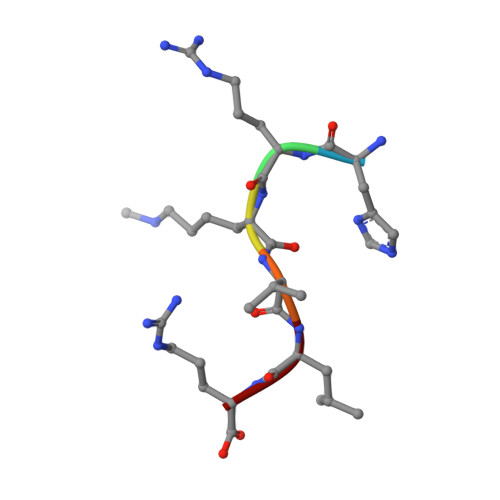
Molecular recognition of histone lysine methylation by the Polycomb group repressor dSfmbt
Grimm, C., Matos, R., Ly-Hartig, N., Steuerwald, U., Lindner, D., Rybin, V., Muller, J., Muller, C.W.(2009) EMBO J 28: 1965-1977
- PubMed: 19494831 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/emboj.2009.147
- Primary Citation Related Structures:
3H6Z - PubMed Abstract:
Polycomb group (PcG) proteins repress transcription by modifying chromatin structure in target genes. dSfmbt is a subunit of the Drosophila melanogaster PcG protein complex PhoRC and contains four malignant brain tumour (MBT) repeats involved in the recognition of various mono- and dimethylated histone peptides. Here, we present the crystal structure of the four-MBT-repeat domain of dSfmbt in complex with a mono-methylated histone H4 peptide. Only a single histone peptide binds to the four-MBT-repeat domain. Mutational analyses show high-affinity binding with low peptide sequence selectivity through combinatorial interaction of the methyl-lysine with an aromatic cage and positively charged flanking residues with the surrounding negatively charged surface of the fourth MBT repeat. dSfmbt directly interacts with the PcG protein Scm, a related MBT-repeat protein with similar methyl-lysine binding activity. dSfmbt and Scm co-occupy Polycomb response elements of target genes in Drosophila and they strongly synergize in the repression of these target genes, suggesting that the combined action of these two MBT proteins is crucial for Polycomb silencing.
- European Molecular Biology Laboratory, Grenoble Outstation, Grenoble, France.
Organizational Affiliation: