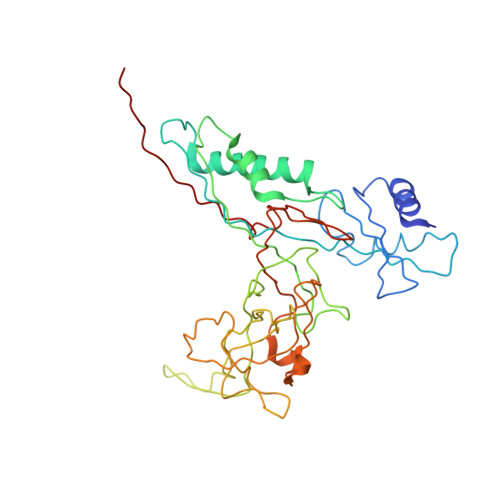
The structure of gene product 6 of bacteriophage T4, the hinge-pin of the baseplate.
Aksyuk, A.A., Leiman, P.G., Shneider, M.M., Mesyanzhinov, V.V., Rossmann, M.G.(2009) Structure 17: 800-808
- PubMed: 19523898 Search on PubMed
- DOI: https://doi.org/10.1016/j.str.2009.04.005
- Primary Citation Related Structures:
3H2T, 3H3W, 3H3Y - PubMed Abstract:
The baseplate of bacteriophage T4 is a multicomponent protein complex, which controls phage attachment to the host. It assembles from six wedges and a central hub. During infection the baseplate undergoes a large conformational change from a dome-shaped to a flat, star-shaped structure. We report the crystal structure of the C-terminal half of gene product (gp) 6 and investigate its motion with respect to the other proteins during the baseplate rearrangement. Six gp6 dimers interdigitate, forming a ring that maintains the integrity of the baseplate in both conformations. One baseplate wedge contains an N-terminal dimer of gp6, whereas neighboring wedges are tied together through the C-terminal dimer of gp6. The dimeric interactions are preserved throughout the rearrangement of the baseplate. However, the hinge angle between the N- and C-terminal parts of gp6 changes by approximately 15 degrees , accounting for a 10 A radial increase in the diameter of the gp6 ring.
- Department of Biological Sciences, Purdue University, 915 W. State Street, West Lafayette, IN 47907-2054, USA.
Organizational Affiliation: