Structure and function of an ADP-ribose-dependent transcriptional regulator of NAD metabolism
Huang, N., De Ingeniis, J., Galeazzi, L., Mancini, C., Korostelev, Y.D., Rakhmaninova, A.B., Gelfand, M.S., Rodionov, D.A., Raffaelli, N., Zhang, H.(2009) Structure 17: 939-951
- PubMed: 19604474 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.str.2009.05.012
- Primary Citation Related Structures:
3GZ5, 3GZ6, 3GZ8 - PubMed Abstract:
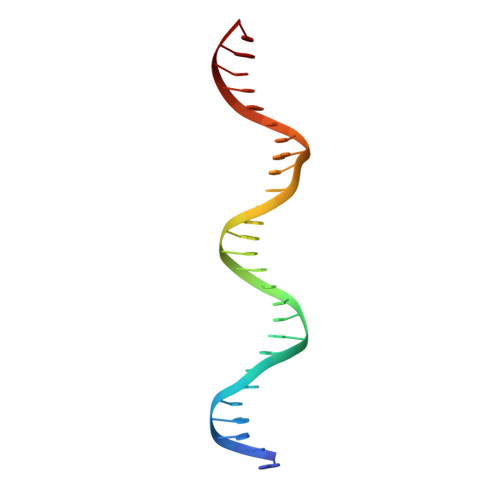
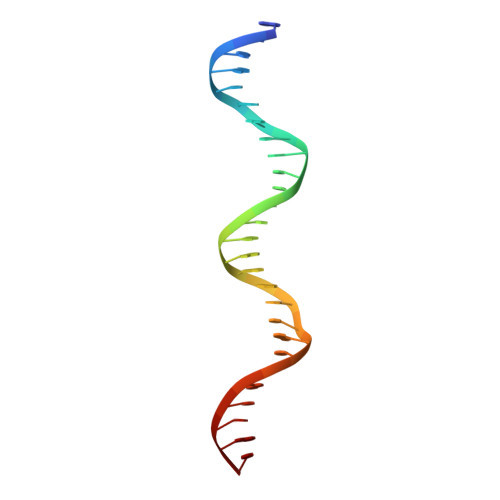
Besides its function as an essential redox cofactor, nicotinamide adenine dinucleotide (NAD) also serves as a consumable substrate for several reactions with broad impact on many cellular processes. NAD homeostasis appears to be tightly controlled, but the mechanism of its regulation is little understood. Here we demonstrate that a previously predicted bacterial transcriptional regulator, NrtR, represses the transcription of NAD biosynthetic genes in vitro. The NAD metabolite ADP-ribose functions as an activator suppressing NrtR repressor activity. The presence of high ADP-ribose levels in the cell is indicative of active NAD turnover in bacteria, which could signal the activation of NAD biosynthetic gene expression via inhibiting the repressor function of NrtR. By comparing the crystal structures of NrtR in complex with DNA and with ADP-ribose, we identified a "Nudix switch" element that likely plays a critical role in the allosteric regulation of DNA binding and repressor function of NrtR.
- Department of Biochemistry, University of Texas Southwestern Medical Center, Dallas, TX 75390, USA.
Organizational Affiliation: