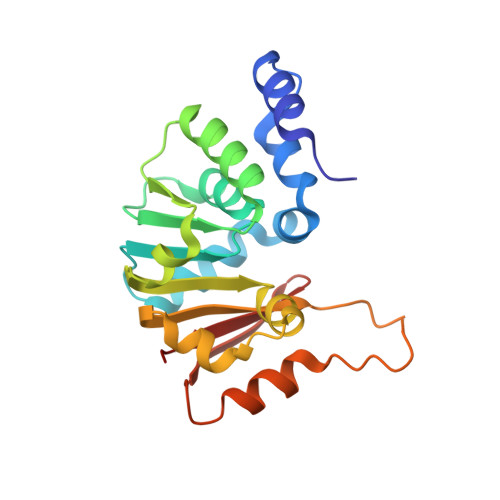
Structural bases for 16 S rRNA methylation catalyzed by ArmA and RmtB methyltransferases
Schmitt, E., Galimand, M., Panvert, M., Courvalin, P., Mechulam, Y.(2009) J Mol Biology 388: 570-582
- PubMed: 19303884 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2009.03.034
- Primary Citation Related Structures:
3FRH, 3FRI, 3FZG - PubMed Abstract:
Aminoglycosides are used extensively for the treatment of severe infections due to Gram-negative bacteria. However, certain species have become highly resistant after acquisition of genes for methyltransferases which catalyze post-transcriptional methylation of N7-G1405 in 16 S rRNA of 30 S ribosomal subunits. Inactivation of this enzymatic activity is therefore an important challenge for development of an effective therapy. The present work describes the crystallographic structures of methyltransferases RmtB and ArmA from clinical isolates. Together with biochemical experiments, the 3D structures indicate that the N-terminal domain specific for this family of methyltransferases is required for enzymatic activity. Site-directed mutagenesis has enabled important residues for catalysis and RNA binding to be identified. These high-resolution structures should underpin the design of potential inhibitors of these enzymes, which could be used to restore the activity of aminoglycosides against resistant pathogens.
- Laboratoire de Biochimie, Ecole Polytechnique, Centre National de la Recherche Scientifique, Palaiseau Cedex, France. emma@botrytis.polytechnique.fr.
Organizational Affiliation: