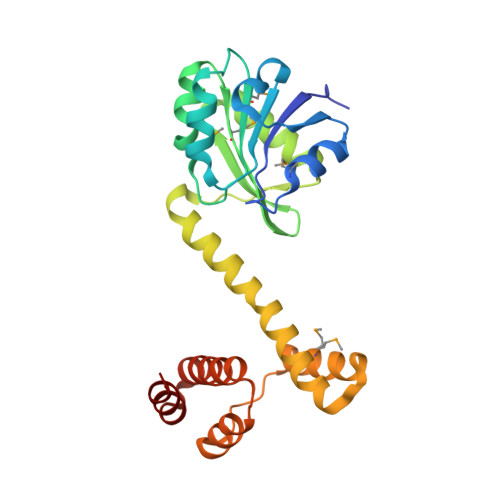
Crystal structure of Protein of Unknown Function from the 6-Phosphogluconate Dehydrogenase-Like Family (NP_601885.1) from CORYNEBACTERIUM GLUTAMICUM ATCC 13032 KITASATO at 2.07 A resolution
Joint Center for Structural Genomics (JCSG)To be published.