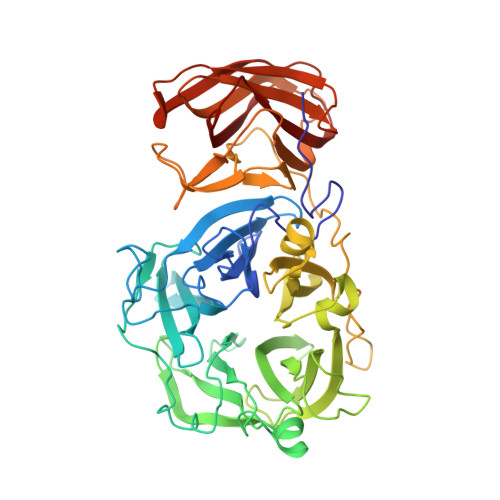
Structural analysis of a glycoside hydrolase family 43 arabinoxylan arabinofuranohydrolase in complex with xylotetraose reveals a different binding mechanism compared with other members of the same family.
Vandermarliere, E., Bourgois, T.M., Winn, M.D., van Campenhout, S., Volckaert, G., Delcour, J.A., Strelkov, S.V., Rabijns, A., Courtin, C.M.(2009) Biochem J 418: 39-47
- PubMed: 18980579 Search on PubMed
- DOI: https://doi.org/10.1042/BJ20081256
- Primary Citation Related Structures:
3C7E, 3C7F, 3C7G, 3C7H, 3C7O - PubMed Abstract:
AXHs (arabinoxylan arabinofuranohydrolases) are alpha-L-arabinofuranosidases that specifically hydrolyse the glycosidic bond between arabinofuranosyl substituents and xylopyranosyl backbone residues of arabinoxylan. Bacillus subtilis was recently shown to produce an AXH that cleaves arabinose units from O-2- or O-3-mono-substituted xylose residues: BsAXH-m2,3 (B. subtilis AXH-m2,3). Crystallographic analysis reveals a two-domain structure for this enzyme: a catalytic domain displaying a five-bladed beta-propeller fold characteristic of GH (glycoside hydrolase) family 43 and a CBM (carbohydrate-binding module) with a beta-sandwich fold belonging to CBM family 6. Binding of substrate to BsAXH-m2,3 is largely based on hydrophobic stacking interactions, which probably allow the positional flexibility needed to hydrolyse both arabinose substituents at the O-2 or O-3 position of the xylose unit. Superposition of the BsAXH-m2,3 structure with known structures of the GH family 43 exo-acting enzymes, beta-xylosidase and alpha-L-arabinanase, each in complex with their substrate, reveals a different orientation of the sugar backbone.
- Laboratory for Biocrystallography, Department of Pharmaceutical Sciences, Katholieke Universiteit Leuven, Herestraat 49, O&N II, bus 822, 3000 Leuven, Belgium.
Organizational Affiliation: