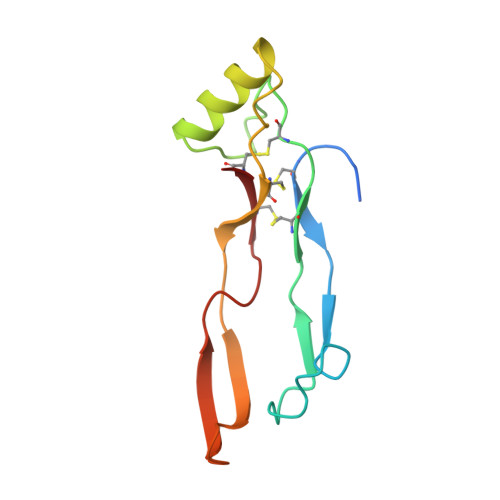
Crystal structure of human bone morphogenetic protein-2 at 2.7 A resolution.
Scheufler, C., Sebald, W., Hulsmeyer, M.(1999) J Mol Biology 287: 103-115
- PubMed: 10074410 Search on PubMed
- DOI: https://doi.org/10.1006/jmbi.1999.2590
- Primary Citation Related Structures:
3BMP - PubMed Abstract:
Homodimeric bone morphogenetic protein-2 (BMP-2) is a member of the transforming growth factor beta (TGF-beta) superfamily that induces bone formation and regeneration, and determines important steps during early stages of embryonic development in vertebrates and non-vertebrates. BMP-2 can interact with two types of receptor chains, as well as with proteins of the extracellular matrix and several regulatory proteins. We report here the crystal structure of human BMP-2 determined by molecular replacement and refined to an R-value of 24.2 % at 2.7 A resolution. A common scaffold of BMP-2, BMP-7 and the TGF-betas, i.e. the cystine-knot motif and two finger-like double-stranded beta-sheets, can be superimposed with r. m.s. deviations of around 1 A. In contrast to the TGF-betas, the structure of BMP-2 shows differences in the flexibility of the N terminus and the orientation of the central alpha-helix as well as two external loops at the fingertips with respect to the scaffold. This is also known from the BMP-7 model. Small secondary structure elements in the loop regions of BMP-2 and BMP-7 seem to be specific for the respective BMP-subgroup. Two identical helix-finger clefts and two distinct cavities located around the central 2-fold axis of the dimer show characteristic shapes, polarity and surface charges. The possible function of these specific features in the interaction of BMP-2 with its binding partners is discussed.
- Physiological Chemistry II, University of Würzburg, Würzburg, 97074, Germany.
Organizational Affiliation: