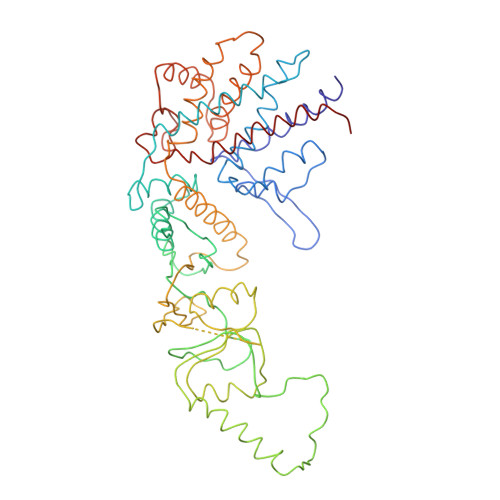
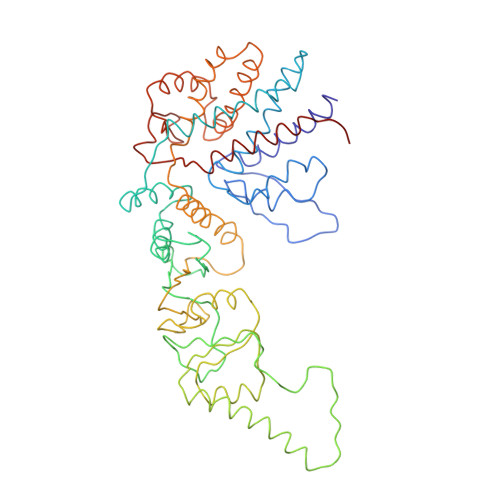
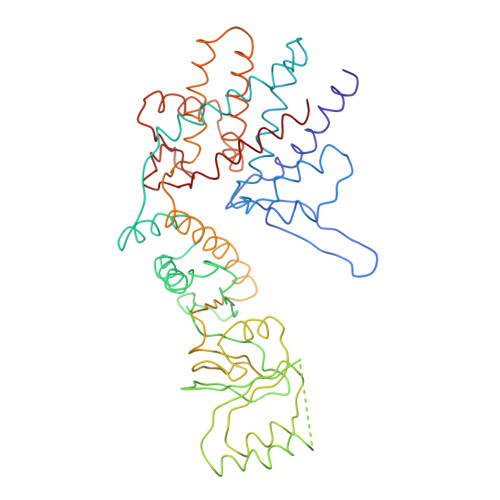
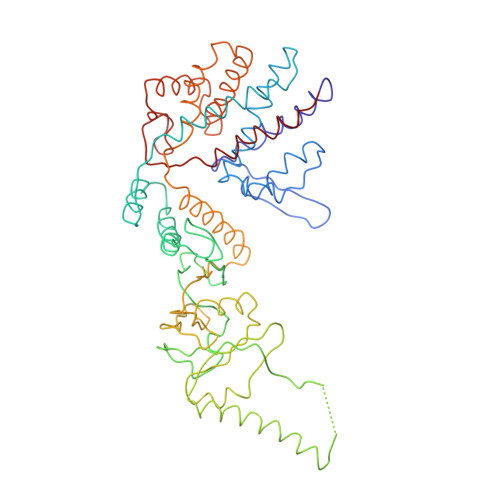
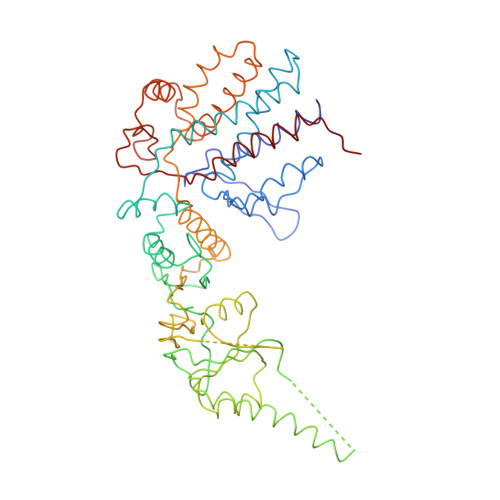
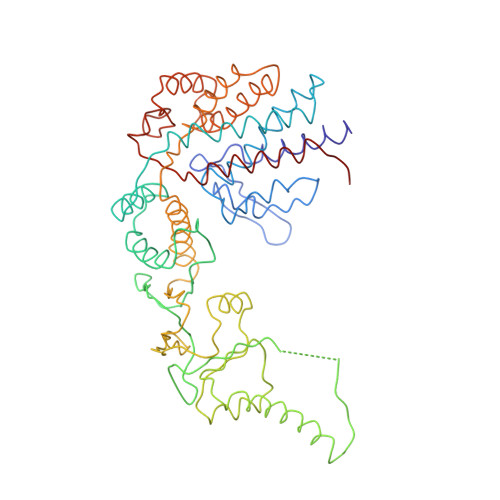
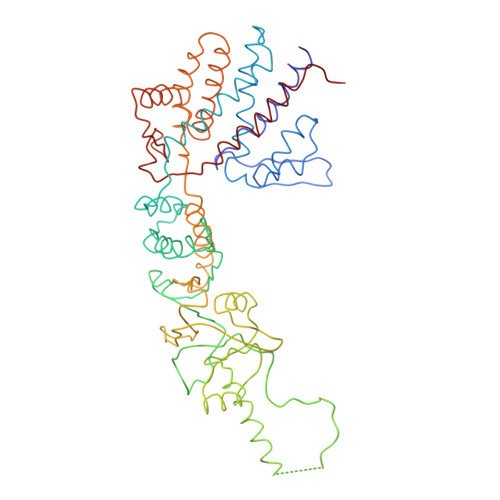
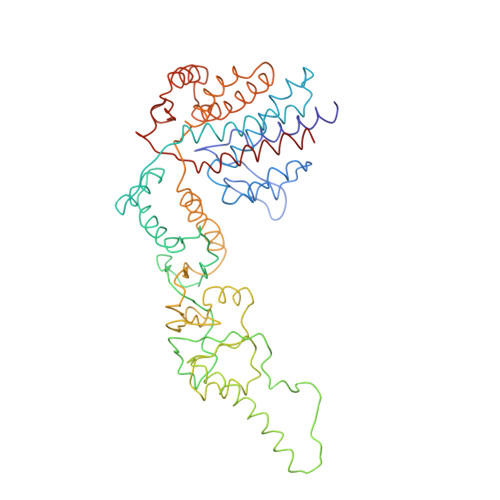
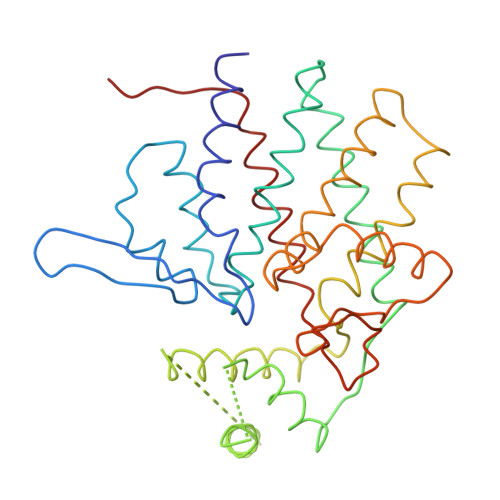
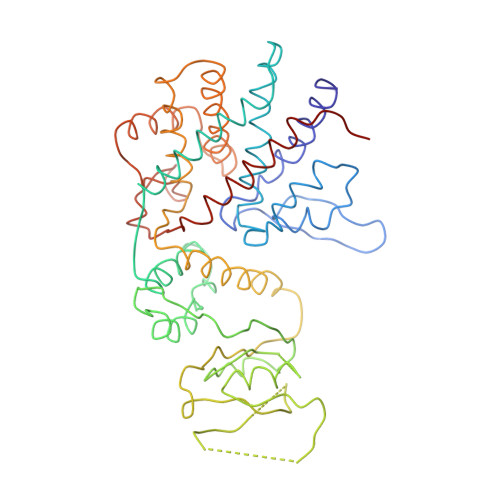
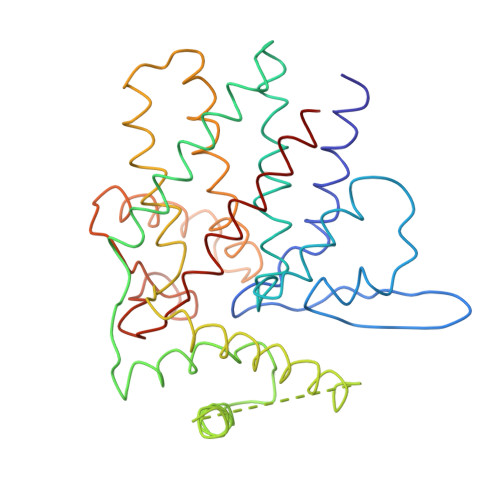
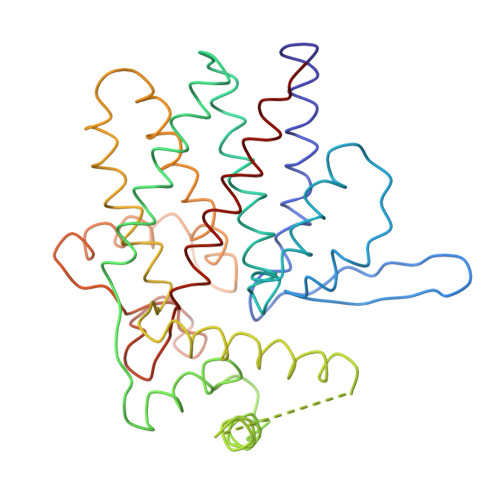
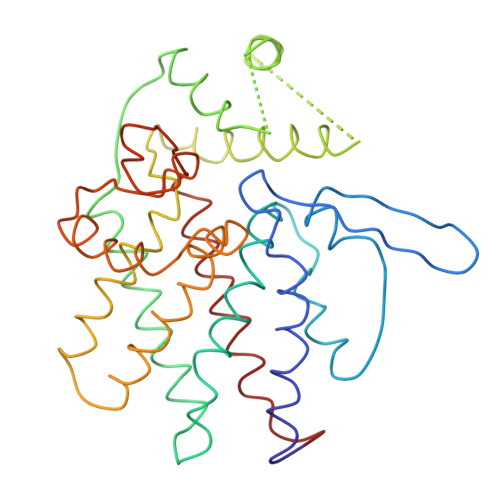
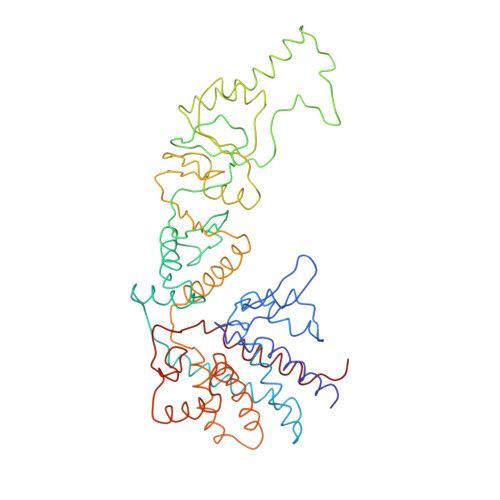
Crystal Structure of the Open Conformation of the Mammalian Chaperonin Cct in Complex with Tubulin.
Munoz, I.G., Yebenes, H., Zhou, M., Mesa, P., Serna, M., Park, A.Y., Bragado-Nilsson, E., Beloso, A., De Carcer, G., Malumbres, M., Robinson, C.V., Valpuesta, J.M., Montoya, G.(2011) Nat Struct Mol Biol 18: 14
- PubMed: 21151115 Search on PubMed
- DOI: https://doi.org/10.1038/nsmb.1971
- Primary Citation Related Structures:
2XSM - PubMed Abstract:
Protein folding is assisted by molecular chaperones. CCT (chaperonin containing TCP-1, or TRiC) is a 1-MDa oligomer that is built by two rings comprising eight different 60-kDa subunits. This chaperonin regulates the folding of important proteins including actin, α-tubulin and β-tubulin. We used an electron density map at 5.5 Å resolution to reconstruct CCT, which showed a substrate in the inner cavities of both rings. Here we present the crystal structure of the open conformation of this nanomachine in complex with tubulin, providing information about the mechanism by which it aids tubulin folding. The structure showed that the substrate interacts with loops in the apical and equatorial domains of CCT. The organization of the ATP-binding pockets suggests that the substrate is stretched inside the cavity. Our data provide the basis for understanding the function of this chaperonin.
- Macromolecular Crystallography Group, Structural Biology and Biocomputing Programme, Spanish National Cancer Research Centre (CNIO), Madrid, Spain.
Organizational Affiliation: