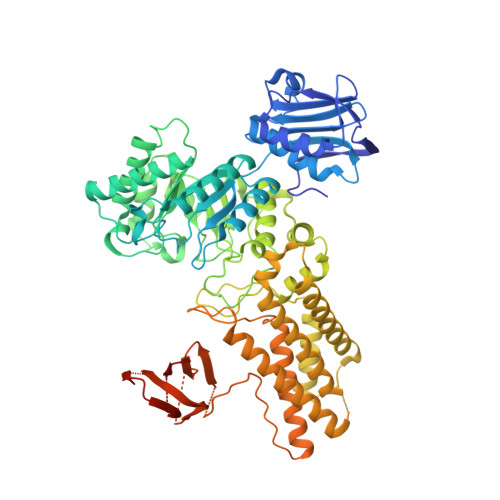
Inhibition of O-Glcnacase Using a Potent and Cell-Permeable Inhibitor Does not Induce Insulin Resistance in 3T3-L1 Adipocytes.
Macauley, M.S., He, Y., Gloster, T.M., Stubbs, K.A., Davies, G.J., Vocadlo, D.J.(2010) Chem Biol 17: 937
- PubMed: 20851343 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.chembiol.2010.07.006
- Primary Citation Related Structures:
2XJ7 - PubMed Abstract:
To probe increased O-GlcNAc levels as an independent mechanism governing insulin resistance in 3T3-L1 adipocytes, a new class of O-GlcNAcase (OGA) inhibitor was studied. 6-Acetamido-6-deoxy-castanospermine (6-Ac-Cas) is a potent inhibitor of OGA. The structure of 6-Ac-Cas bound in the active site of an OGA homolog reveals structural features contributing to its potency. Treatment of 3T3-L1 adipocytes with 6-Ac-Cas increases O-GlcNAc levels in a dose-dependent manner. These increases in O-GlcNAc levels do not induce insulin resistance functionally, measured using a 2-deoxyglucose (2-DOG) uptake assay, or at the molecular level, determined by evaluating levels of phosphorylated IRS-1 and Akt. These results, and others described, provide a structural blueprint for improved inhibitors and collectively suggest that increased O-GlcNAc levels, brought about by inhibition of OGA, does not by itself cause insulin resistance in 3T3-L1 adipocytes.
- Department of Chemistry, Simon Fraser University, Burnaby, BC, Canada.
Organizational Affiliation: