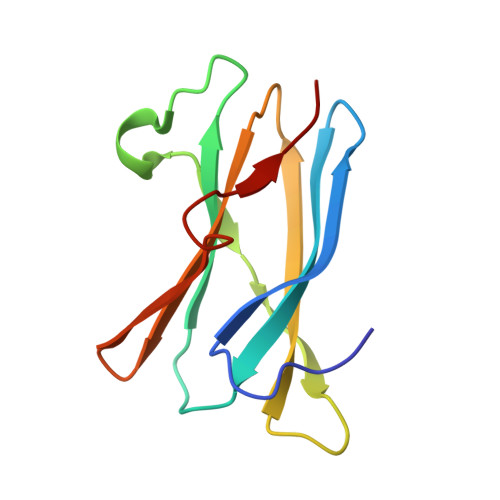
Crystal Structures of the Starch-Binding Domain from Rhizopus Oryzae Glucoamylase Reveal a Polysaccharide-Binding Path.
Tung, J.-Y., Chang, M.D., Chou, W., Liu, Y.-Y., Yeh, Y., Chang, F., Lin, S., Qiu, Z., Sun, Y.-J.(2008) Biochem J 416: 27
- PubMed: 18588504 Search on PubMed
- DOI: https://doi.org/10.1042/BJ20080580
- Primary Citation Related Structures:
2V8L, 2V8M, 2VQ4 - PubMed Abstract:
GA (glucoamylase) hydrolyses starch and polysaccharides to beta-D-glucose. RoGA (Rhizopus oryzae GA) consists of two functional domains, an N-terminal SBD (starch-binding domain) and a C-terminal catalytic domain, which are connected by an O-glycosylated linker. In the present study, the crystal structures of the SBD from RoGA (RoGACBM21) and the complexes with beta-cyclodextrin (SBD-betaCD) and maltoheptaose (SBD-G7) were determined. Two carbohydrate binding sites, I (Trp(47)) and II (Tyr(32)), were resolved and their binding was co-operative. Besides the hydrophobic interaction, two unique polyN loops comprising consecutive asparagine residues also participate in the sugar binding. A conformational change in Tyr(32) was observed between unliganded and liganded SBDs. To elucidate the mechanism of polysaccharide binding, a number of mutants were constructed and characterized by a quantitative binding isotherm and Scatchard analysis. A possible binding path for long-chain polysaccharides in RoGACBM21 was proposed.
- Institute of Bioinformatics and Structural Biology, National Tsing Hua University, Hsinchu 30013, Taiwan.
Organizational Affiliation: