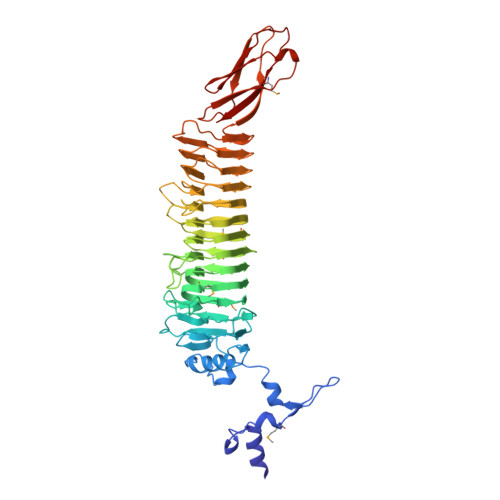
An intersubunit active site between supercoiled parallel beta helices in the trimeric tailspike endorhamnosidase of Shigella flexneri Phage Sf6.
Muller, J.J., Barbirz, S., Heinle, K., Freiberg, A., Seckler, R., Heinemann, U.(2008) Structure 16: 766-775
- PubMed: 18462681 Search on PubMed
- DOI: https://doi.org/10.1016/j.str.2008.01.019
- Primary Citation Related Structures:
2VBE, 2VBK, 2VBM - PubMed Abstract:
Sf6 belongs to the Podoviridae family of temperate bacteriophages that infect gram-negative bacteria by insertion of their double-stranded DNA. They attach to their hosts specifically via their tailspike proteins. The 1.25 A crystal structure of Shigella phage Sf6 tailspike protein (Sf6 TSP) reveals a conserved architecture with a central, right-handed beta helix. In the trimer of Sf6 TSP, the parallel beta helices form a left-handed, coiled-beta coil with a pitch of 340 A. The C-terminal domain consists of a beta sandwich reminiscent of viral capsid proteins. Further crystallographic and biochemical analyses show a Shigella cell wall O-antigen fragment to bind to an endorhamnosidase active site located between two beta-helix subunits each anchoring one catalytic carboxylate. The functionally and structurally related bacteriophage, P22 TSP, lacks sequence identity with Sf6 TSP and has its active sites on single subunits. Sf6 TSP may serve as an example for the evolution of different host specificities on a similar general architecture.
- Max-Delbrück-Centrum für Molekulare Medizin, Robert-Rössle-Str. 10, 13125 Berlin, Germany.
Organizational Affiliation: