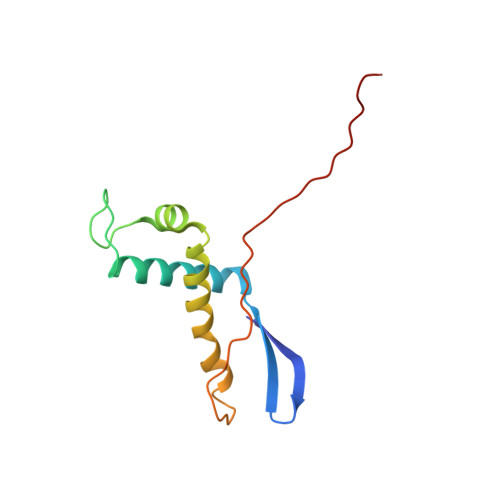
Structures of and interactions between domains of trigger factor from Thermotoga maritima.
Martinez-Hackert, E., Hendrickson, W.A.(2007) Acta Crystallogr D Biol Crystallogr 63: 536-547
- PubMed: 17372359 Search on PubMed
- DOI: https://doi.org/10.1107/S090744490700964X
- Primary Citation Related Structures:
2NSA, 2NSB, 2NSC - PubMed Abstract:
Trigger factor (TF) is a eubacterial chaperone that associates with ribosomes at the peptide-exit tunnel and also occurs in excess free in the cytosol. TF is a three-domain protein that appears to exist in a dynamic equilibrium of oligomerization states and interdomain conformations. X-ray crystallography and chemical cross-linking were used to study the roles of the N- and C-terminal domains of Thermotoga maritima TF in TF oligomerization and chaperone activity. The structural conservation of both the N- and C-terminal TF domains was unambiguously established. The biochemical and crystallographic data reveal a tendency for these domains to partake in diverse and apparently nonspecific protein-protein interactions. It is found that the T. maritima and Escherichia coli TF surfaces lack evident exposed hydrophobic patches. Taken together, these data suggest that TF chaperones could interact with nascent proteins via hydrophilic surfaces.
- Department of Biochemistry and Molecular Biophysics, Columbia University, New York, NY 10032, USA.
Organizational Affiliation: