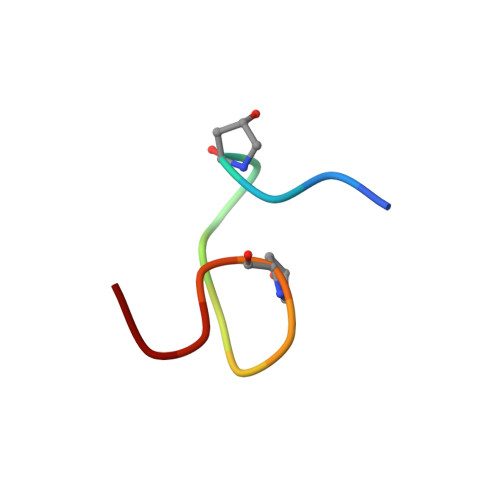
Solution NMR studies of the plant peptide hormone CEP inform function.
Bobay, B.G., Digennaro, P., Scholl, E., Imin, N., Djordjevic, M.A., McK Bird, D.(2013) FEBS Lett 587: 3979-3985
- PubMed: 24211833 Search on PubMed
- DOI: https://doi.org/10.1016/j.febslet.2013.10.033
- Primary Citation Related Structures:
2MFM, 2MFO - PubMed Abstract:
The C-terminally Encoded Peptide (CEP) family of regulatory peptides controls root development in vascular plants. Here, we present the first NMR structures of CEP. We show that root-knot nematode (RKN: Meloidogyne spp.) also encodes CEP, presumably to mimic plant CEP as part of their stereotypic, parasitic interaction with vascular plants. Molecular dynamics simulations of plant- and nematode-encoded CEP displaying known posttranslational modifications (PTM) provided insight into the structural effects of PTM and the conformational plasticity and rigidity of CEP. Potential mechanisms of action are discussed with respect to the structure and sampling of conformational space.
- Department of Molecular and Structural Biochemistry, NC State University, Raleigh, NC 27695, United States.
Organizational Affiliation: