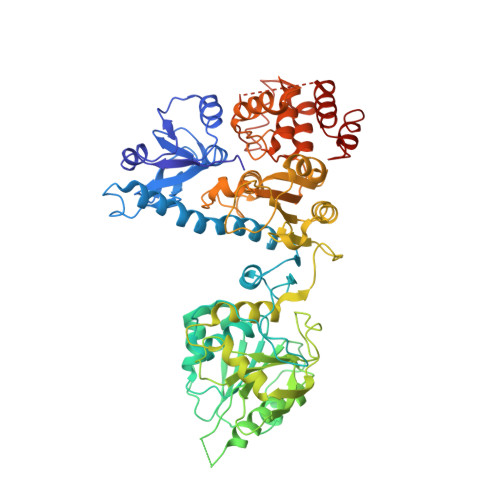
Bluetongue Virus Vp4 is an RNA-Capping Assembly Line.
Sutton, G., Grimes, J.M., Stuart, D.I., Roy, P.(2007) Nat Struct Mol Biol 14: 449
- PubMed: 17417654 Search on PubMed
- DOI: https://doi.org/10.1038/nsmb1225
- Primary Citation Related Structures:
2JH8, 2JH9, 2JHA, 2JHC, 2JHP - PubMed Abstract:
Eukaryotic organisms cap the 5' ends of their messenger RNAs by a series of four chemical reactions. Some viruses achieve this using a single molecule; the crystal structure of such an enzyme from bluetongue virus reveals an elongated modular architecture that provides a scaffold for an assemblage of active sites, two contributed by a domain of novel structure.
- Division of Structural Biology, The Wellcome Trust Centre for Human Genetics, University of Oxford, Roosevelt Drive, Headington, Oxford, OX3 7BN, UK.
Organizational Affiliation: