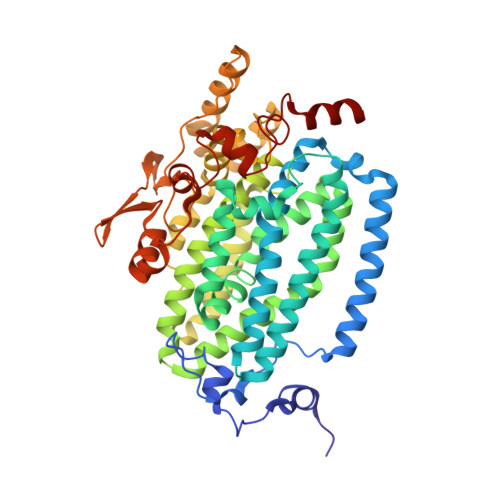
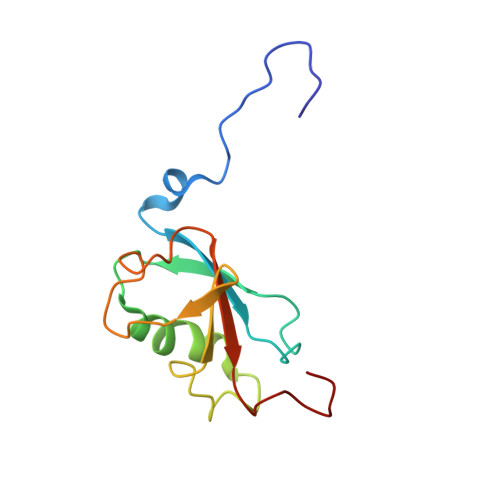
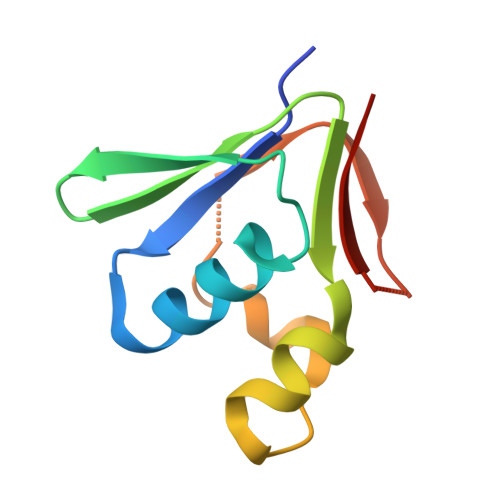
X-ray Structure of a Hydroxylase-Regulatory Protein Complex from a Hydrocarbon-Oxidizing Multicomponent Monooxygenase, Pseudomonas sp. OX1 Phenol Hydroxylase.
Sazinsky, M.H., Dunten, P.W., McCormick, M.S., Didonato, A., Lippard, S.J.(2006) Biochemistry 45: 15392-15404
- PubMed: 17176061 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/bi0618969
- Primary Citation Related Structures:
2INN, 2INP - PubMed Abstract:
Phenol hydroxylase (PH) belongs to a family of bacterial multicomponent monooxygenases (BMMs) with carboxylate-bridged diiron active sites. Included are toluene/o-xylene (ToMO) and soluble methane (sMMO) monooxygenase. PH hydroxylates aromatic compounds, but unlike sMMO, it cannot oxidize alkanes despite having a similar dinuclear iron active site. Important for activity is formation of a complex between the hydroxylase and a regulatory protein component. To address how structural features of BMM hydroxylases and their component complexes may facilitate the catalytic mechanism and choice of substrate, we determined X-ray structures of native and SeMet forms of the PH hydroxylase (PHH) in complex with its regulatory protein (PHM) to 2.3 A resolution. PHM binds in a canyon on one side of the (alphabetagamma)2 PHH dimer, contacting alpha-subunit helices A, E, and F approximately 12 A above the diiron core. The structure of the dinuclear iron center in PHH resembles that of mixed-valent MMOH, suggesting an Fe(II)Fe(III) oxidation state. Helix E, which comprises part of the iron-coordinating four-helix bundle, has more pi-helical character than analogous E helices in MMOH and ToMOH lacking a bound regulatory protein. Consequently, conserved active site Thr and Asn residues translocate to the protein surface, and an approximately 6 A pore opens through the four-helix bundle. Of likely functional significance is a specific hydrogen bond formed between this Asn residue and a conserved Ser side chain on PHM. The PHM protein covers a putative docking site on PHH for the PH reductase, which transfers electrons to the PHH diiron center prior to O2 activation, suggesting that the regulatory component may function to block undesired reduction of oxygenated intermediates during the catalytic cycle. A series of hydrophobic cavities through the PHH alpha-subunit, analogous to those in MMOH, may facilitate movement of the substrate to and/or product from the active site pocket. Comparisons between the ToMOH and PHH structures provide insights into their substrate regiospecificities.
- Department of Chemistry, Massachusetts Institute of Technology, Cambridge, Massachusetts 02139, USA.
Organizational Affiliation: