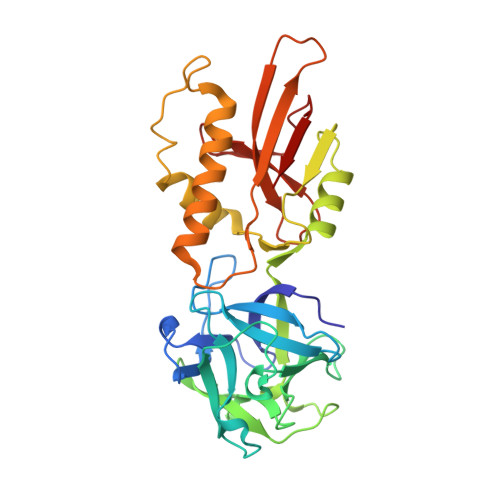
Crystal structure of the marasmius oreades mushroom lectin in complex with a xenotransplantation epitope.
Grahn, E., Askarieh, G., Holmner, A., Tateno, H., Winter, H.C., Goldstein, I.J., Krengel, U.(2007) J Mol Biology 369: 710-721
- PubMed: 17442345 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2007.03.016
- Primary Citation Related Structures:
2IHO - PubMed Abstract:
MOA, a lectin from the mushroom Marasmius oreades, is one of the few reagents that specifically agglutinate blood group B erythrocytes. Further, it is the only lectin known to have exclusive specificity for Galalpha(1,3)Gal-containing sugar epitopes, which are antigens that pose a severe barrier to animal-to-human organ transplantation. We describe here the structure of MOA at 2.4 A resolution, in complex with the linear trisaccharide Galalpha(1,3)Galbeta(1,4)GlcNAc. The structure is dimeric, with two distinct domains per protomer: the N-terminal lectin module adopts a ricinB/beta-trefoil fold and contains three putative carbohydrate-binding sites, while the C-terminal domain serves as a dimerization interface. This latter domain, which has an unknown function, reveals a novel fold with intriguing conservation of an active site cleft. A number of indications suggest that MOA may have an enzymatic function in addition to the sugar-binding properties.
- Department of Chemistry, University of Oslo, P.O. Box 1033, Blindern, NO-0315 Oslo, Norway. elin.grahn@kjemi.uio.no <elin.grahn@kjemi.uio.no>
Organizational Affiliation: