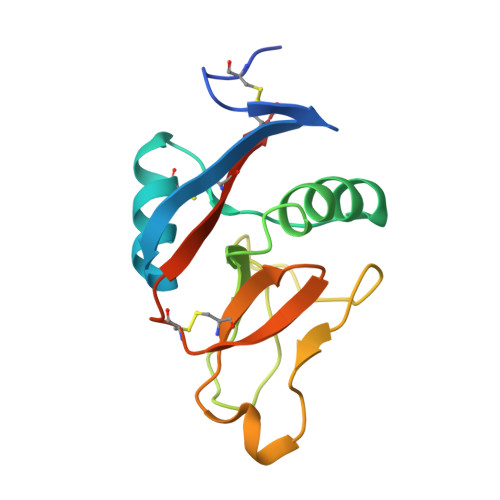
Structure of the Fungal Beta-Glucan-Binding Immune Receptor Dectin-1: Implications for Function.
Brown, J., O'Callaghan, C.A., Marshall, A.S.J., Gilbert, R.J.C., Siebold, C., Gordon, S., Brown, G.D., Jones, E.Y.(2007) Protein Sci 16: 1042
- PubMed: 17473009 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1110/ps.072791207
- Primary Citation Related Structures:
2BPD, 2BPE, 2BPH, 2CL8 - PubMed Abstract:
The murine molecule dectin-1 (known as the beta-glucan receptor in humans) is an immune cell surface receptor implicated in the immunological defense against fungal pathogens. Sequence analysis has indicated that the dectin-1 extracellular domain is a C-type lectin-like domain, and functional studies have established that it binds fungal beta-glucans. We report several dectin-1 crystal structures, including a high-resolution structure and a 2.8 angstroms resolution structure in which a short soaked natural beta-glucan is trapped in the crystal lattice. In vitro characterization of dectin-1 in the presence of its natural ligand indicates higher-order complex formation between dectin-1 and beta-glucans. These combined structural and biophysical data considerably extend the current knowledge of dectin-1 structure and function, and suggest potential mechanisms of defense against fungal pathogens.
- CR-UK Receptor Structure Research Group, Division of Structural Biology, Wellcome Trust Centre for Human Genetics, University of Oxford, Oxford, OX3 7BN, United Kingdom.
Organizational Affiliation: