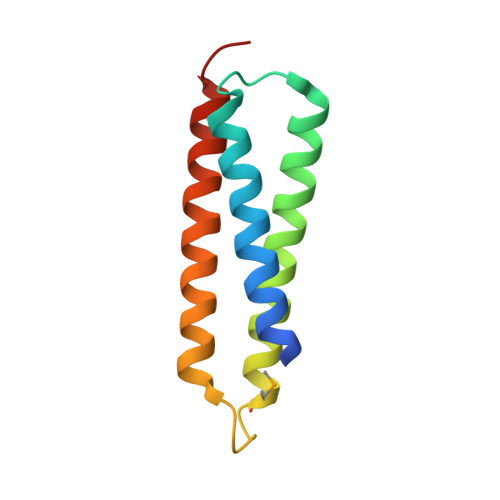
Structural Analysis of the Interaction between the Snare Tlg1 and Vps51.
Fridmann-Sirkis, Y., Kent, H.M., Lewis, M.J., Evans, P.R., Pelham, H.R.B.(2006) Traffic 7: 182
- PubMed: 16420526 Search on PubMed
- DOI: https://doi.org/10.1111/j.1600-0854.2005.00374.x
- Primary Citation Related Structures:
2C5I, 2C5J, 2C5K - PubMed Abstract:
Membrane fusion in cells involves the interaction of SNARE proteins on apposing membranes. Formation of SNARE complexes is preceded by tethering events, and a number of protein complexes that are thought to mediate this have been identified. The VFT or GARP complex is required for endosome-Golgi traffic in yeast. It consists of four subunits, one of which, Vps51, has been shown to bind specifically to the SNARE Tlg1, which participates in the same fusion event. We have determined the structure of the N-terminal domain of Tlg1 bound to a peptide from the N terminus of Vps51. Binding depends mainly on residues 18-30 of Vps51. These form a short helix which lies in a conserved groove in the three-helix bundle formed by Tlg1. Surprisingly, although both Vps51 and Tlg1 are required for transport to the late Golgi from endosomes, removal of the Tlg1-binding sequences from Vps51 does not block such traffic in vivo. Thus, this particular interaction cannot be crucial to the process of vesicle docking or fusion.
- MRC Laboratory of Molecular Biology, Hills Road, Cambridge CB2 2QH, UK.
Organizational Affiliation: