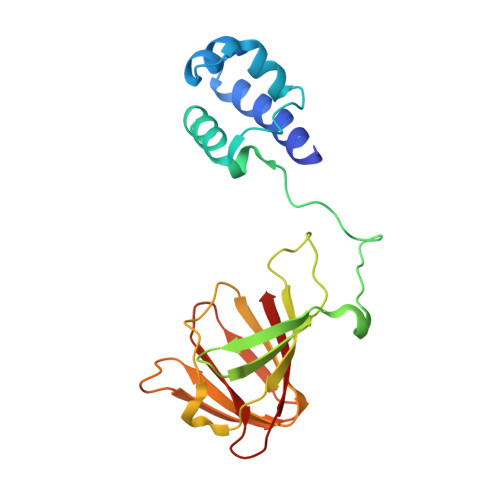
Structure and Reactivity of Hydroxypropylphosphonic Acid Epoxidase in Fosfomycin Biosynthesis by a Cation- and Flavin-Dependent Mechanism.
Mcluskey, K., Cameron, S., Hammerschmidt, F., Hunter, W.N.(2005) Proc Natl Acad Sci U S A 102: 14221
- PubMed: 16186494 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.0504314102
- Primary Citation Related Structures:
2BNM, 2BNN, 2BNO - PubMed Abstract:
The biosynthesis of fosfomycin, an oxirane antibiotic in clinical use, involves a unique epoxidation catalyzed by (S)-2-hydroxypropylphosphonic acid epoxidase (HPPE). The reaction is essentially dehydrogenation of a secondary alcohol. A high-resolution crystallographic analysis reveals that the HPPE subunit displays a two-domain combination. The C-terminal or catalytic domain has the cupin fold that binds a divalent cation, whereas the N-terminal domain carries a helix-turn-helix motif with putative DNA-binding helices positioned 34 A apart. The structure of HPPE serves as a model for numerous proteins, of ill-defined function, predicted to be transcription factors but carrying a cupin domain at the C terminus. Structure-reactivity analyses reveal conformational changes near the catalytic center driven by the presence or absence of ligand, that HPPE is a Zn(2+)/Fe(2+)-dependent epoxidase, proof that flavin mononucleotide is required for catalysis, and allow us to propose a simple mechanism that is compatible with previous experimental data. The participation of the redox inert Zn(2+) in the mechanism is surprising and indicates that Lewis acid properties of the metal ions are sufficient to polarize the substrate and, aided by flavin mononucleotide reduction, facilitate the epoxidation.
- Division of Biological Chemistry and Molecular Microbiology, School of Life Sciences, University of Dundee, Dundee DD1 5EH, United Kingdom.
Organizational Affiliation: