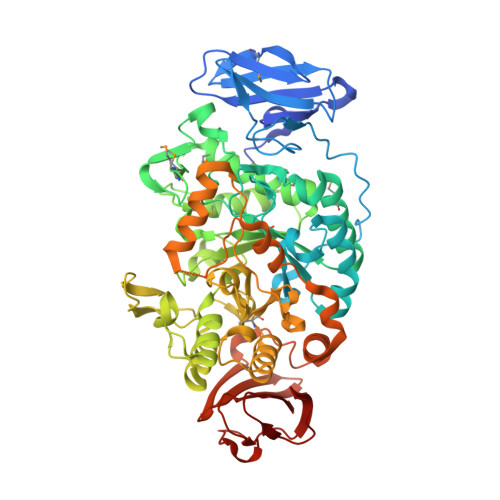
Crystal Structure of Maltooligosyltrehalose Trehalohydrolase from Deinococcus Radiodurans in Complex with Disaccharides
Timmins, J., Leiros, H.-K.S., Leonard, G., Leiros, I., Mcsweeney, S.(2005) J Mol Biology 347: 949
- PubMed: 15784255 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2005.02.011
- Primary Citation Related Structures:
2BHU, 2BHY, 2BHZ - PubMed Abstract:
Trehalose (alpha-D-glucopyranosyl-1,1-alpha-D-glucopyranose) is a non-reducing diglucoside found in various organisms that serves as a carbohydrate reserve and as an agent that protects against a variety of physical and chemical stresses. Deinococcus radiodurans possesses an alternative biosynthesis pathway for the synthesis of trehalose from maltooligosaccharides. This reaction is mediated by two enzymes: maltooligosyltrehalose synthase (MTSase) and maltooligosyltrehalose trehalohydrolase (MTHase). Here, we present the 1.1A resolution crystal structure of MTHase. It consists of three major domains: two beta-sheet domains and a conserved glycosidase (beta/alpha)8 barrel catalytic domain. Three subdomains consisting of short insertions were identified within the catalytic domain. Subsequently, structures of MTHase in complex with maltose and trehalose were obtained at 1.2 A and 1.5 A resolution, respectively. These structures reveal the importance of the three inserted subdomains in providing the key residues required for substrate recognition. Trehalose is recognised specifically in the +1 and +2 binding subsites by an extensive hydrogen-bonding network and a strong hydrophobic stacking interaction in between two aromatic residues. Moreover, upon binding to maltose, which mimics the substrate sugar chain, a major concerted conformational change traps the sugar chain in the active site. The presence of magnesium in the active site of the MTHase-maltose complex suggests that MTHase activity may be regulated by divalent cations.
- Macromolecular Crystallography Group, European Synchrotron Radiation Facility, B.P. 220, 6 rue Jules Horowitz, F-38043 Grenoble Cedex, France.
Organizational Affiliation: