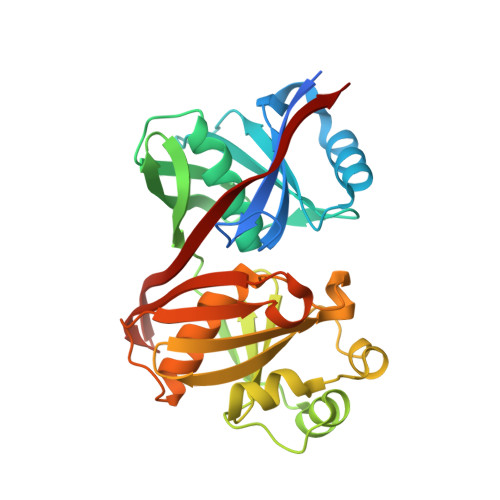
Structure and function of the phenazine biosynthetic protein PhzF from Pseudomonas fluorescens
Blankenfeldt, W., Kuzin, A.P., Skarina, T., Korniyenko, Y., Tong, L., Bayer, P., Janning, P., Thomashow, L.S., Mavrodi, D.V.(2004) Proc Natl Acad Sci U S A 101: 16431-16437
- PubMed: 15545603 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.0407371101
- Primary Citation Related Structures:
1SDJ, 1U1V, 1U1W, 1U1X, 1XUA, 1XUB - PubMed Abstract:
Phenazines produced by Pseudomonas and Streptomyces spp. are heterocyclic nitrogen-containing metabolites with antibiotic, antitumor, and antiparasitic activity. The antibiotic properties of pyocyanin, produced by Pseudomonas aeruginosa, were recognized in the 1890s, although this blue phenazine is now known to be a virulence factor in human disease. Despite their biological significance, the biosynthesis of phenazines is not fully understood. Here we present structural and functional studies of PhzF, an enzyme essential for phenazine synthesis in Pseudomonas spp. PhzF shares topology with diaminopimelate epimerase DapF but lacks the same catalytic residues. The structure of PhzF in complex with its substrate, trans-2,3-dihydro-3-hydroxyanthranilic acid, suggests that it is an isomerase using the conserved glutamate E45 to abstract a proton from C3 of the substrate. The proton is returned to C1 of the substrate after rearrangement of the double-bond system, yielding an enol that converts to the corresponding ketone. PhzF is a dimer that may be bifunctional, providing a shielded cavity for ketone dimerization via double Schiff-base formation to produce the phenazine scaffold. Our proposed mechanism is supported by mass and NMR spectroscopy. The results are discussed in the context of related structures and protein sequences of unknown biochemical function.
- Max Planck Institute of Molecular Physiology, Otto-Hahn-Strasse 11, 44227 Dortmund, Germany.
Organizational Affiliation: