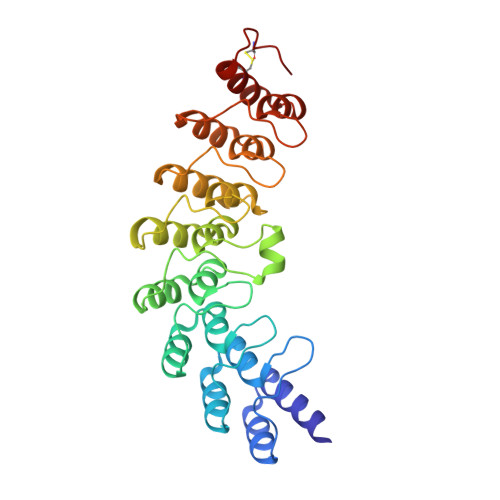
Structural basis for recognition of 2',5'-linked oligoadenylates by human ribonuclease L
Tanaka, N., Nakanishi, M., Kusakabe, Y., Goto, Y., Kitade, Y., Nakamura, K.T.(2004) EMBO J 23: 3929-3938
- PubMed: 15385955 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/sj.emboj.7600420
- Primary Citation Related Structures:
1WDY - PubMed Abstract:
An interferon-induced endoribonuclease, ribonuclease L (RNase L), is implicated in both the molecular mechanism of action of interferon and the fundamental control of RNA stability in mammalian cells. RNase L is catalytically active only after binding to an unusual activator molecule containing a 5'-phosphorylated 2',5'-linked oligoadenylate (2-5A), in the N-terminal half. Here, we report the crystal structure of the N-terminal ankyrin repeat domain (ANK) of human RNase L complexed with the activator 2-5A. This is the first structural view of an ankyrin repeat structure directly interacting with a nucleic acid, rather than with a protein. The ANK domain folds into eight ankyrin repeat elements and forms an extended curved structure with a concave surface. The 2-5A molecule is accommodated at a concave site and directly interacts with ankyrin repeats 2-4. Interestingly, two structurally equivalent 2-5A binding motifs are found at repeats 2 and 4. The structural basis for 2-5A recognition by ANK is essential for designing stable 2-5As with a high likelihood of activating RNase L.
- School of Pharmaceutical Sciences, Showa University, Hatanodai, Shinagawa-ku, Tokyo, Japan. ntanaka@pharm.showa-u.ac.jp
Organizational Affiliation: