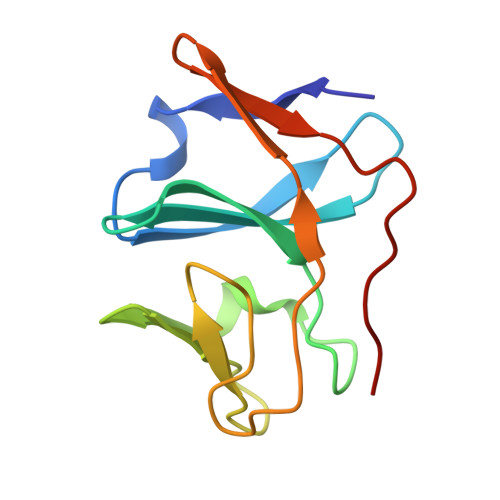
Structure of T4moC, the Rieske-type ferredoxin component of toluene 4-monooxygenase.
Moe, L.A., Bingman, C.A., Wesenberg, G.E., Phillips, G.N., Fox, B.G.(2006) Acta Crystallogr D Biol Crystallogr 62: 476-482
- PubMed: 16627939 Search on PubMed
- DOI: https://doi.org/10.1107/S0907444906006056
- Primary Citation Related Structures:
1VM9 - PubMed Abstract:
The structure of the Rieske-type ferredoxin (T4moC) from toluene 4-monooxygenase was determined by X-ray crystallography in the [2Fe-2S](2+) state at a resolution of 1.48 A using single-wavelength anomalous dispersion phasing with the [2Fe-2S] center. The structure consists of ten beta-strands arranged into the three antiparallel beta-sheet topology observed in all Rieske proteins. Trp69 of T4moC is adjacent to the [2Fe-2S] centre, which displaces a loop containing the conserved Pro81 by approximately 8 A away from the [2Fe-2S] cluster compared with the Pro loop in the closest structural and functional homolog, the Rieske-type ferredoxin BphF from biphenyl dioxygenase. In addition, T4moC contains five hydrogen bonds to the [2Fe-2S] cluster compared with three hydrogen bonds in BphF. Moreover, the electrostatic surface of T4moC is distinct from that of BphF. These structural differences are identified as possible contributors to the evolutionary specialization of soluble Rieske-type ferredoxins between the diiron monooxygenases and cis-dihydrodiol-forming dioxygenases.
- Department of Biochemistry, University of Wisconsin-Madison, USA.
Organizational Affiliation: