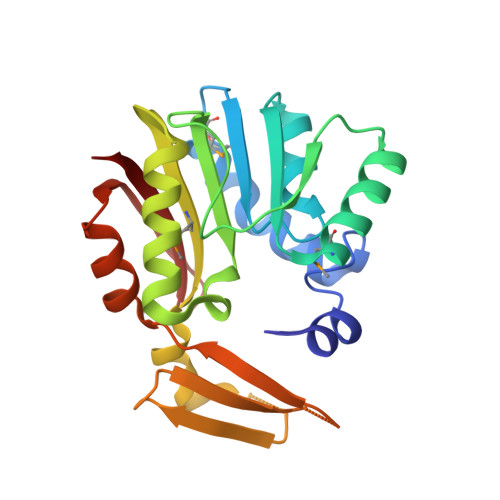
Crystal structure of SAM-dependent methyltransferase from Pyrococcus horikoshii.
Pampa, K.J., Madan Kumar, S., Hema, M.K., Kumara, K., Naveen, S., Kunishima, N., Lokanath, N.K.(2017) Acta Crystallogr F Struct Biol Commun 73: 706-712
- PubMed: 29199993 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S2053230X17016648
- Primary Citation Related Structures:
1VE3 - PubMed Abstract:
Methyltransferases (MTs) are enzymes involved in methylation that are needed to perform cellular processes such as biosynthesis, metabolism, gene expression, protein trafficking and signal transduction. The cofactor S-adenosyl-L-methionine (SAM) is used for catalysis by SAM-dependent methyltransferases (SAM-MTs). The crystal structure of Pyrococcus horikoshii SAM-MT was determined to a resolution of 2.1 Å using X-ray diffraction. The monomeric structure consists of a Rossmann-like fold (domain I) and a substrate-binding domain (domain II). The cofactor (SAM) molecule binds at the interface between adjacent subunits, presumably near to the active site(s) of the enzyme. The observed dimeric state might be important for the catalytic function of the enzyme.
- Department of Studies in Biotechnology, University of Mysore, Manasagangotri, Mysuru, Karnataka 570 006, India.
Organizational Affiliation: