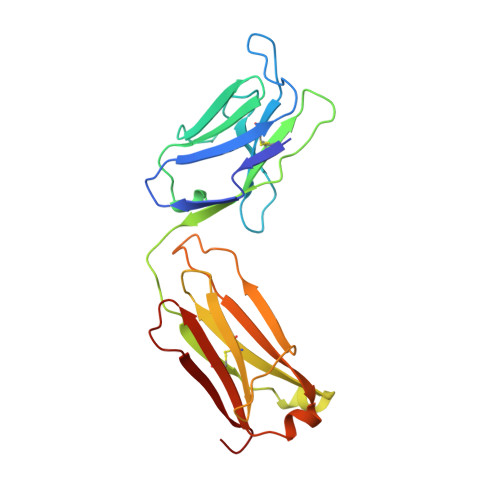
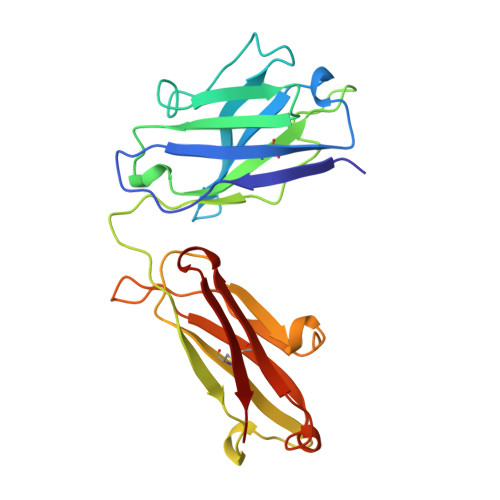
X-ray structure of the uncomplexed anti-tumor antibody BR96 and comparison with its antigen-bound form.
Sheriff, S., Chang, C.Y., Jeffrey, P.D., Bajorath, J.(1996) J Mol Biology 259: 938-946
- PubMed: 8683596 Search on PubMed
- DOI: https://doi.org/10.1006/jmbi.1996.0371
- Primary Citation Related Structures:
1UCB - PubMed Abstract:
The X-ray structure of the uncomplexed human chimeric Fab' of the anti-tumor antibody BR96 has been determined at 2.6 A resolution. The structure has been compared with Lewis Y antigen-complexed structures of BR96 which were determined previously. The comparison reveals segmental motions and/or conformational rearrangements of three CDR loops (L1, L3, and H2), whereas CDR H3 does not undergo changes upon complexation despite its significant main-chain contacts to the carbohydrate antigen. In light of the uncomplexed chimeric Fab' structure reported here, the previously observed high mobility of the CL:CH1 domains of the complexed chimeric BR96 Fab is rationalized as a "swinging" motion approximately about the axis of the elbow bend.
- Bristol-Myers Squibb Pharmaceutical Research Institute, Princeton, NJ 08543-4000, USA.
Organizational Affiliation: