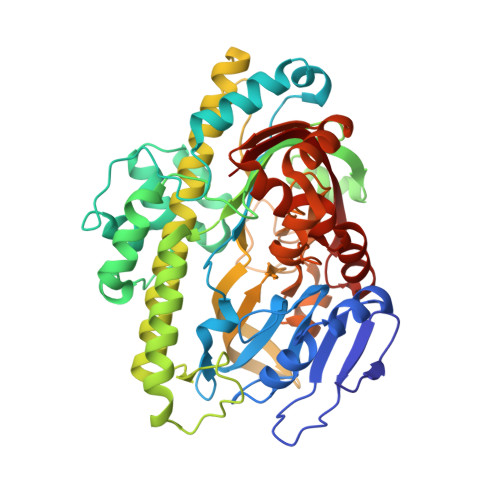
Crystal structure of an RtcB homolog protein (PH1602-extein protein) from Pyrococcus horikoshii reveals a novel fold
Okada, C., Maegawa, Y., Yao, M., Tanaka, I.(2006) Proteins 63: 1119-1122
- PubMed: 16485279 Search on PubMed
- DOI: https://doi.org/10.1002/prot.20912
- Primary Citation Related Structures:
1UC2 - Division of Biological Sciences, Graduate School of Science, Hokkaido University, Sapporo, Japan.
Organizational Affiliation: