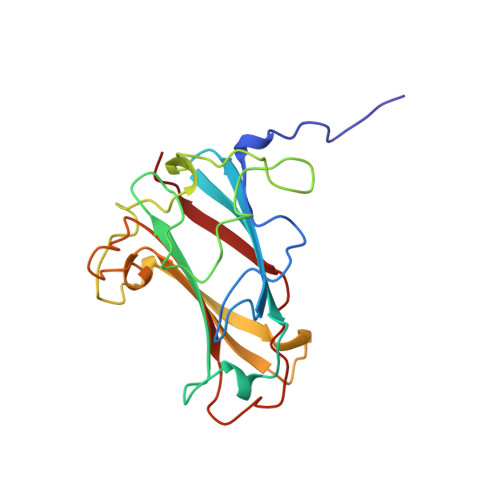
Structure of the human adenovirus serotype 2 fiber head domain at 1.5 A resolution.
van Raaij, M.J., Louis, N., Chroboczek, J., Cusack, S.(1999) Virology 262: 333-343
- PubMed: 10502512 Search on PubMed
- DOI: https://doi.org/10.1006/viro.1999.9849
- Primary Citation Related Structures:
1QHV - PubMed Abstract:
Adenovirus binds to its receptor via the head domain of its fiber protein. We have crystallized the adenovirus serotype 2 (subgroup C) receptor binding domain and solved the structure at 1.5 A resolution by the molecular replacement technique using the known adenovirus type 5 head structure. Included in the high-resolution model are 306 water molecules, five alternative side chain conformations, and individual anisotropic temperature factors for each atom. The overall structure of the serotype 2 head is very similar to its serotype 5 homologue, apart from differences in some of the flexible loops. All but subgroup B adenoviruses are believed to use the recently identified protein CAR (Coxsackievirus and adenovirus receptor) as receptor. By comparison of the two structures and sequence alignment of CAR binding and non-CAR binding serotype fiber heads, we discuss possible receptor binding sites and propose a receptor binding site in a crevice between two monomers on the side of the trimer. The structural basis of the extraordinary stability of the fiber head trimer is also discussed.
- c/o Institut Laue Langevin, 6 rue Jules Horowitz, Grenoble Cedex 9, 38000, France.
Organizational Affiliation: