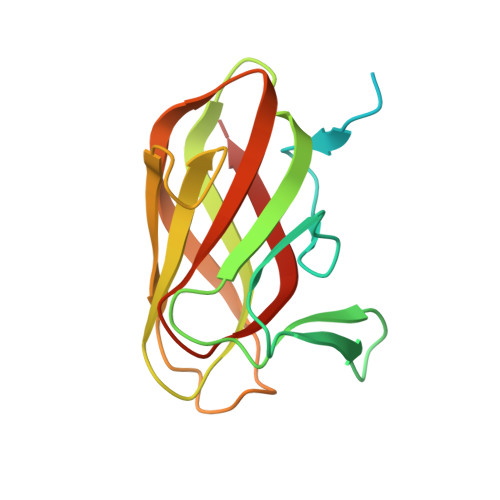
Structure and ligand binding of carbohydrate-binding module CsCBM6-3 reveals similarities with fucose-specific lectins and "galactose-binding" domains
Boraston, A.B., Notenboom, V., Warren, R.A.J., Kilburn, D.G., Rose, D.R., Davies, G.(2003) J Mol Biology 327: 659-669
- PubMed: 12634060 Search on PubMed
- DOI: https://doi.org/10.1016/s0022-2836(03)00152-9
- Primary Citation Related Structures:
1NAE, 1O8P, 1O8S, 1OD3 - PubMed Abstract:
Carbohydrate-binding polypeptides, including carbohydrate-binding modules (CBMs) from polysaccharidases, and lectins, are widespread in nature. Whilst CBMs are classically considered distinct from lectins, in that they are found appended to polysaccharide-degrading enzymes, this distinction is blurring. The crystal structure of CsCBM6-3, a "sequence-family 6" CBM in a xylanase from Clostridium stercorarium, at 2.3 A reveals a similar, all beta-sheet fold to that from MvX56, a module found in a family 33 glycoside hydrolase sialidase from Micromonospora viridifaciens, and the lectin AAA from Anguilla anguilla. Sequence analysis leads to the classification of MvX56 and AAA into a family distinct from that containing CsCBM6-3. Whilst these polypeptides are similar in structure they have quite different carbohydrate-binding specificities. AAA is known to bind fucose; CsCBM6-3 binds cellulose, xylan and other beta-glucans. Here we demonstrate that MvX56 binds galactose, lactose and sialic acid. Crystal structures of CsCBM6-3 in complex with xylotriose, cellobiose, and laminaribiose, 2.0 A, 1.35 A, and 1.0 A resolution, respectively, reveal that the binding site of CsCBM6-3 resides on the same polypeptide face as for MvX56 and AAA. Subtle differences in the ligand-binding surface give rise to the different specificities and biological activities, further blurring the distinction between classical lectins and CBMs.
- Department of Chemistry, The University of York, Heslington, York, YO10 5YW, UK.
Organizational Affiliation: