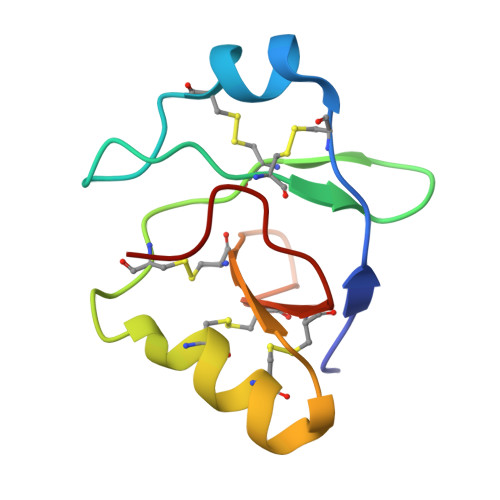
Necrosis-Suppressing Effector Protein ChEC88 Adopts a Novel Structural Motif Conserved Among Genus-Spanning Hemibiotrophic Phytopathogens.
Ohki, S., Takahara, H., Imamura, T., Sakane, K., Bai, A., Sasaki, K., Nishiuchi, T., Mori, M.(2025) Plants (Basel) 14
- PubMed: 40872185
- DOI: https://doi.org/10.3390/plants14162562
- Primary Citation of Related Structures:
9UB0 - PubMed Abstract:
Phytopathogenic fungi secrete numerous effector proteins to disrupt plant defenses. At present, their sequence-structure-function relationships remain poorly understood owing to their diversity. Comprehensive understanding of conserved effectors is necessary to elucidate the molecular relationship between fungi and plants. To fill this research gap, we investigated the Colletotrichum higginsianum effector candidate (ChEC)-88 specifically expressed during infection. Notably, similar to the biotrophy-associated secreted protein 3 (BAS3) from Pyricularia oryzae , ChEC88 inhibited plant cell death caused by necrosis- and ethylene-inducing peptide 1-like protein (NLP1). Nuclear magnetic resonance analysis results revealed that ChEC88 adopted a novel pseudo two-fold symmetrical three-dimensional structure. Homology modeling suggested that BAS3 exhibited a ChEC88-like conformation despite sharing less than 50% sequence identity. Through PSI-BLAST searches, we found that ChEC88 homologs were conserved in various hemibiotrophic phytopathogenic fungi, including Colletotrichum , P. oryzae , and Fusarium species. Functional assays demonstrated that all of the representative homologs suppressed NLP1-induced plant cell death. Mutation experiments identified the residues critical for ChEC88 function. Overall, our findings suggest that hemibiotrophic phytopathogenic fungi share a conserved immune-suppression strategy mediated by ChEC88-like proteins and that such effectors possibly originated from a common ancestral lineage of phytopathogenic fungi.
- Center for Nano Materials and Technology (CNMT), Japan Advanced Institute of Science and Technology (JAIST), Nomi 923-1292, Ishikawa, Japan.
Organizational Affiliation: