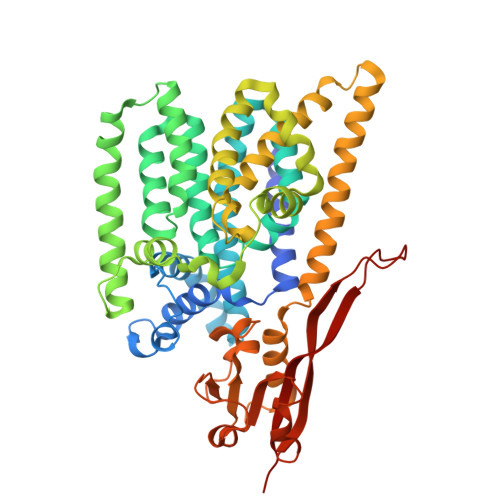
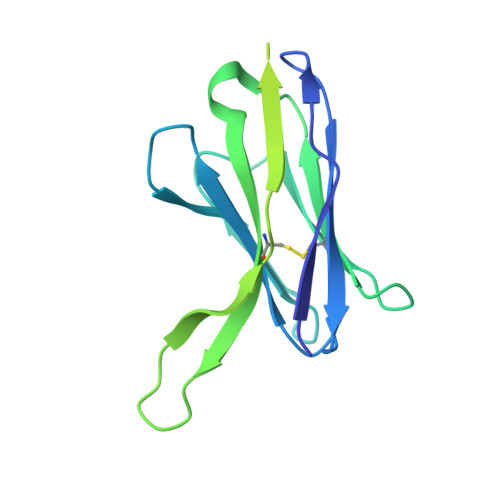
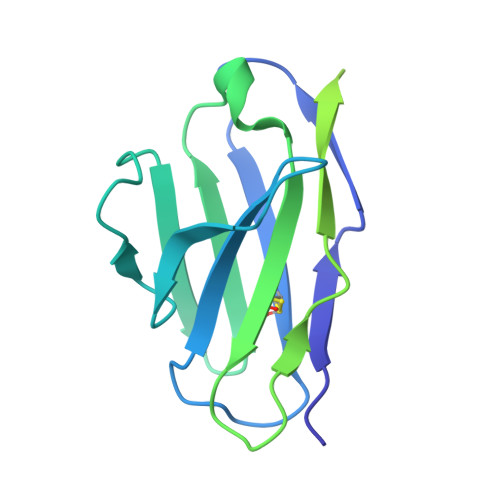
Structures of ALG3/9/12 reveal the assembly logic of the N-glycan oligomannose core.
Alexander, J.A.N., Chen, S.Y., Mukherjee, S., de Capitani, M., Irobalieva, R.N., Rossi, L., Agrawal, P., Kowal, J., Meirelles, M.A., Aebi, M., Reymond, J.L., Kossiakoff, A.A., Riniker, S., Locher, K.P.(2026) Nat Chem Biol
- PubMed: 41807832 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41589-026-02164-7
- Primary Citation Related Structures:
9S6R, 9S6S, 9S6T, 9S6U - PubMed Abstract:
Asparagine-linked glycans are essential for the maturation and function of most eukaryotic secretory proteins. The biosynthesis and transfer of dolichylpyrophosphate-anchored GlcNAc 2 Man 9 Glc 3 glycan is a highly conserved process occurring in the endoplasmic reticulum (ER) membrane and involving over a dozen membrane proteins whose dysfunction is linked to congenital disorders of glycosylation (CDGs). Three membrane-integral mannosyltransferases, ALG3, ALG9 and ALG12, mediate four consecutive mannosylation reactions that convert GlcNAc 2 Man 5 to GlcNAc 2 Man 9 . Here, using chemoenzymatically synthesized lipid-linked glycan donor and acceptor analogs, we recapitulated this biosynthetic pathway in vitro. High-resolution cryo-electron microscopy structures of pseudo-Michaelis complexes of each step revealed how the branched glycan is accurately synthesized and unwanted side products are averted. Molecular dynamics simulations and mutagenesis studies uncovered a subtle but precise mechanism selecting the dolichylphosphomannose donor substrate over dolichylphosphoglucose, which is also present in the ER membrane. Our results also provide mechanistic explanations for enzyme dysfunction in CDGs and offer opportunities for N-glycan engineering.
- Institute of Molecular Biology and Biophysics, Eidgenössische Technische Hochschule (ETH) Zürich, Zurich, Switzerland.
Organizational Affiliation: