Shared ligand-blocking mechanism but distinct conformational modulation by alpha 5-targeting antibodies BIIG2 and MINT1526A.
Nguyen, A., Heim, J.B., Cordara, G., Chan, M.C., Johannesen, H., Charlesworth, C., Li, M., Azumaya, C.M., Madden, B., Krengel, U., Meves, A., Campbell, M.G.(2025) bioRxiv
- PubMed: 39829743 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1101/2025.01.08.631572
- Primary Citation Related Structures:
8R38, 9B9J, 9B9K - PubMed Abstract:
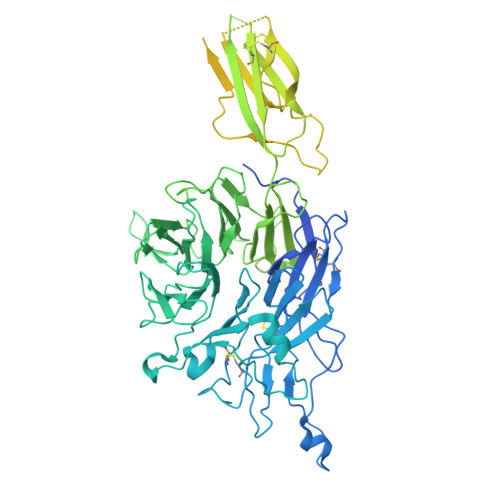
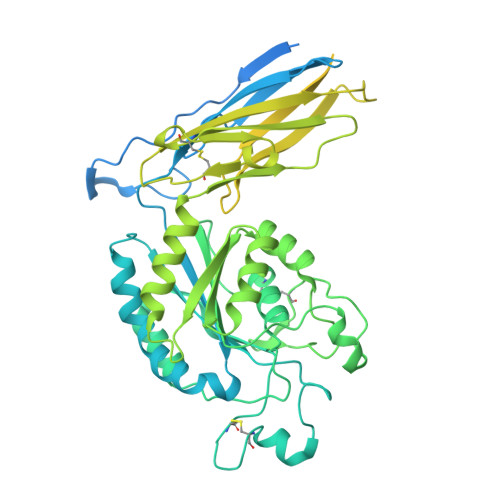
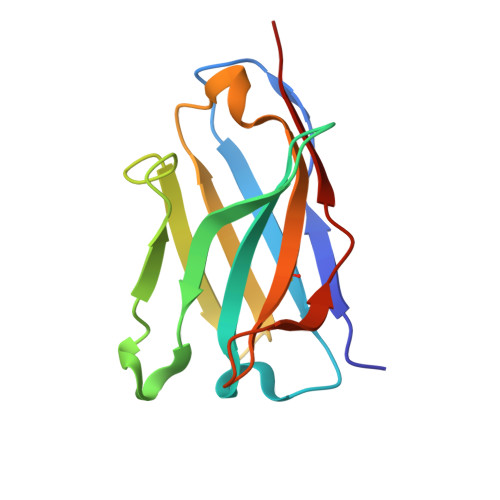
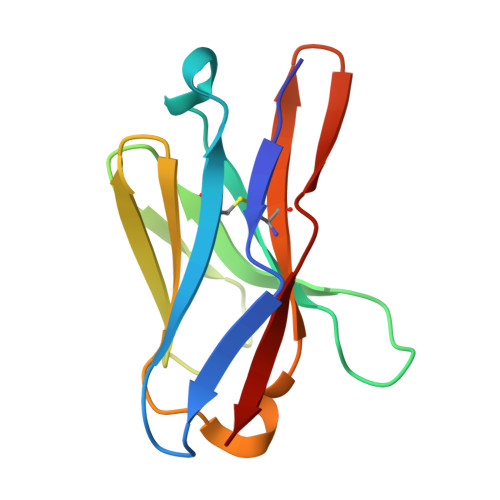
Integrins are a large family of heterodimeric receptors important for cell adhesion and signaling. Integrin α5β1, also known as the fibronectin receptor, is a key mediator of angiogenesis and its dysregulation is associated with tumor proliferation, progression, and metastasis. Despite numerous efforts, α5β1-targeting therapeutics have been unsuccessful in large part due to efficacy and off-target effects. To mediate activation and signaling, integrins undergo drastic conformational changes. However, how therapeutics influence or are affected by integrin conformation remains incompletely characterized. Using cell biology, biophysics, and electron microscopy, we shed light on these relationships by characterizing two potentially therapeutic anti-α5β1 antibodies, BIIG2 and MINT1526A. We show that both antibodies bind α5β1 with nanomolar affinity and reduce angiogenesis in vitro . We demonstrate BIIG2 reduces tumor growth in two human xenograft mouse models and exhibits a strong specificity for connective tissue-resident fibroblasts and melanoma cells. Using electron microscopy, we map out the molecular interfaces mediating the integrin-antibody interactions and reveal that although both antibodies have overlapping epitopes and block fibronectin binding via steric hindrance, the effect on the conformational equilibrium is drastically different. While MINT1526A constricts α5β1's range of flexibility, BIIG2 binds without restricting the available conformational states. These mechanistic insights, coupled with the functional analysis, guide which aspects should be prioritized to avoid off-target effects or partial agonism in the design of future integrin-targeted therapeutics.
- Basic Sciences Division, Fred Hutchinson Cancer Center, Seattle, Washington 98109, USA.
Organizational Affiliation: